Abstract
Background
AT-rich interactive domain 3A (ARID3A) is a member of the ARID family of DNA-binding proteins. Previous reports have shown that ARID3A controls cell growth in a p53-dependent manner. Recently, it has been reported that expression of the ARID3A protein was markedly increased in colon cancer tissue compared with matched normal colonic mucosa. However, little is currently known about the role of ARID3A in colorectal cancer (CRC). The aim of this study was to investigate the clinical significance of ARID3A expression in CRC.
Methods
We examined ARID3A expression in 690 CRC patients using tissue microarray and immunohistochemistry. Kaplan–Meier analysis and Cox proportional hazard models were used to estimate the impact of ARID3A expression on overall survival.
Results
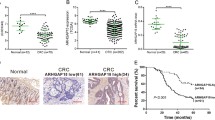
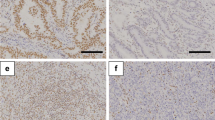
Of the 690 cases, 195 tumors were strongly positive for ARID3A, 187 were weakly positive, and 308 were negative. ARID3A expression in CRC was significantly correlated with age, degree of differentiation, depth of invasion, lymph node metastasis, distant metastasis, tumor–node–metastasis stage, status of microsatellite instability, and carcinoembryonic antigen levels. The overall survival of CRC patients with strong ARID3A expression was significantly longer than that of patients with weak or negative ARID3A expression. We also performed an additional survival analysis on 388 colon cancer patients and 302 rectal cancer patients. In doing so, a favorable prognostic effect of ARID3A expression was revealed only in the colon cancer group (p = 0.002), not in rectal cancer. Moreover, we showed that the effect of ARID3A on the survival was correlated with p53 status. Using multivariate analysis, we found that strong expression of ARID3A was an independent predictor for better prognosis in CRC.
Conclusions
Our data suggested that strong expression of ARID3A may predict a good prognosis in patients with CRC.



Similar content being viewed by others
REFERENCES
Kang UB, Yeom J, Kim HJ, Kim H, Lee C. Expression profiling of more than 3500 proteins of MSS-type colorectal cancer by stable isotope labeling and mass spectrometry. J Proteomics. 2012;75:3050–62.
Wilsker D, Patsialou A, Dallas PB, Moran E. ARID proteins: a diverse family of DNA binding proteins implicated in the control of cell growth, differentiation, and development. Cell Growth Differ. 2002;13:95–106.
Wilsker D, Probst L, Wain HM, Maltais L, Tucker PW, Moran E. Nomenclature of the ARID family of DNA-binding proteins. Genomics. 2005;86:242–51.
Herrscher RF, Kaplan MH, Lelsz DL, Das C, Scheuermann R, Tucker PW. The immunoglobulin heavy-chain matrix-associating regions are bound by Bright: a B cell-specific trans-activator that describes a new DNA-binding protein family. Genes Dev. 1995;9:3067–82.
Rajaiya J, Nixon JC, Ayers N, Desgranges ZP, Roy AL, Webb CF. Induction of immunoglobulin heavy-chain transcription through the transcription factor Bright requires TFII-I. Mol Cell Biol. 2006;26:4758–68.
Webb CF, Smith EA, Medina KL, Buchanan KL, Smithson G, Dou S. Expression of bright at two distinct stages of B lymphocyte development. J Immunol. 1998;160:4747–54.
Kortschak RD, Reimann H, Zimmer M, Eyre HJ, Saint R, Jenne DE. The human dead ringer/bright homolog, DRIL1: cDNA cloning, gene structure, and mapping to D19S886, a marker on 19p13.3 that is strictly linked to the Peutz–Jeghers syndrome. Genomics. 1998;51:288–92.
Nixon JC, Rajaiya JB, Ayers N, Evetts S, Webb CF. The transcription factor, Bright, is not expressed in all human B lymphocyte subpopulations. Cell Immunol. 2004;228:42–53.
Suzuki M, Okuyama S, Okamoto S, Shirasuna K, Nakajima T, Hachiya T, et al. A novel E2F binding protein with Myc-type HLH motif stimulates E2F-dependent transcription by forming a heterodimer. Oncogene. 1998;17:853–65.
Fukuyo Y, Mogi K, Tsunematsu Y, Nakajima T. E2FBP1/hDril1 modulates cell growth through downregulation of promyelocytic leukemia bodies. Cell Death Differ. 2004;11:747–59.
Peeper DS, Shvarts A, Brummelkamp T, Douma S, Koh EY, Daley GQ, et al. A functional screen identifies hDRIL1 as an oncogene that rescues RAS-induced senescence. Nat Cell Biol. 2002;4:148–53.
Ma K, Araki K, Ichwan SJ, Suganuma T, Tamamori-Adachi M, Ikeda MA. E2FBP1/DRIL1, an AT-rich interaction domain-family transcription factor, is regulated by p53. Mol Cancer Res. 2003;1:438–44.
Lestari W, Ichwan SJ, Otsu M, Yamada S, Iseki S, Shimizu S, et al. Cooperation between ARID3A and p53 in the transcriptional activation of p21WAF1 in response to DNA damage. Biochem Biophys Res Commun. 2012;417:710–6.
Wang B, Xiao Z, Ren EC. Redefining the p53 response element. Proc Natl Acad Sci USA. 2009;106:14373–8.
Hilska M, Collan YU, O Laine VJ, Kössi J, Hirsimaki P, Laato M, et al. The significance of tumor markers for proliferation and apoptosis in predicting survival in colorectal cancer. Dis Colon Rectum. 2005;48:2197–208.
Watanabe T, Wu TT, Catalano PJ, Ueki T, Satriano R, Haller DG, et al. Molecular predictors of survival after adjuvant chemotherapy for colon cancer. N Engl J Med. 2001;344:1196–206.
Elsaleh H, Powell B, Soontrapornchai P, Joseph D, Goria F, Spry N, et al. p53 gene mutation, microsatellite instability and adjuvant chemotherapy: impact on survival of 388 patients with Dukes’ C colon carcinoma. Oncology. 2000;58:52–9.
Chen MF, Lee KD, Yeh CH, Chen WC, Huang WS, Chin CC, et al. Role of peroxiredoxin I in rectal cancer and related to p53 status. Int J Radiat Oncol Biol Phys. 2010;78:868–78.
Satow R, Shitashige M, Jigami T, Fukami K, Honda K, Kitabayashi I, et al. β-catenin inhibits promyelocytic leukemia protein tumor suppressor function in colorectal cancer cells. Gastroenterology. 2012;142:572–81.
Cabrera CM, Jimenez P, Concha A, Garrido F, Ruiz-Cabello F. Promyelocytic leukemia (PML) nuclear bodies are disorganized in colorectal tumors with total loss of major histocompatibility complex class I expression and LMP7 downregulation. Tissue Antigens. 2004;63:446–52.
Gurrieri C, Capodieci P, Bernardi R, Scaglioni PP, Nafa K, Rush LJ, et al. Loss of the tumor suppressor PML in human cancers of multiple histologic origins. J Natl Cancer Inst. 2004;96:269–79.
Webb CF, Bryant J, Popowski M, Allred L, Kim D, Harriss J, et al. The ARID family transcription factor bright is required for both hematopoietic stem cell and B lineage development. Mol Cell Biol. 2011;31:1041–53.
An G, Miner CA, Nixon JC, Kincade PW, Bryant J, Tucker PW, et al. Loss of Bright/ARID3a function promotes developmental plasticity. Stem Cells. 2010;28:1560–7.
Ben-Porath I, Thomson MW, Carey VJ, Ge R, Bell GW, Regev A, Weinberg RA. An embryonic stem cell-like gene expression signature in poorly differentiated aggressive human tumors. Nat Genet. 2008;40:499–507.
Vaiopoulos AG, Kostakis ID, Koutsilieris M, Papavassiliou AG. Colorectal cancer stem cells. Stem Cells. 2012;30:363–71.
Saiki Y, Ishimaru S, Mimori K, Takatsuno Y, Nagahara M, Ishii H, et al. Comprehensive analysis of the clinical significance of inducing pluripotent stemness-related gene expression in colorectal cancer cells. Ann Surg Oncol. 2009;16:2638–44.
Saigusa S, Tanaka K, Toiyama Y, Yokoe T, Okugawa Y, Ioue Y, et al. Correlation of CD133, OCT4, and SOX2 in rectal cancer and their association with distant recurrence after chemoradiotherapy. Ann Surg Oncol. 2009;16:3488–98.
Acknowledgment
This work was supported by the Mid-career Researcher Program (2012R1A2A2A01005196) through the National Research Foundation of Korea (NRF) funded by the Ministry of Education, Science and Technology and by a grant of the Korean Health 21 R&D Project, Ministry of Health and Welfare, Republic of Korea (A111218-CP01).
Conflict of interest
No conflicts of interest are declared.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Song, M., Kim, H., Kim, W.K. et al. High Expression of AT-Rich Interactive Domain 3A (ARID3A) is Associated with Good Prognosis in Colorectal Carcinoma. Ann Surg Oncol 21 (Suppl 4), 481–489 (2014). https://doi.org/10.1245/s10434-013-3435-2
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1245/s10434-013-3435-2