-
PDF
- Split View
-
Views
-
Cite
Cite
Veronica Ferrucci, Pasqualino de Antonellis, Francesco Paolo Pennino, Fatemeh Asadzadeh, Antonella Virgilio, Donatella Montanaro, Aldo Galeone, Iolanda Boffa, Ida Pisano, Iolanda Scognamiglio, Luigi Navas, Donatella Diana, Emilia Pedone, Sara Gargiulo, Matteo Gramanzini, Arturo Brunetti, Laura Danielson, Marianeve Carotenuto, Lucia Liguori, Antonio Verrico, Lucia Quaglietta, Maria Elena Errico, Valentina Del Monaco, Valeria D’Argenio, Felice Tirone, Angela Mastronuzzi, Vittoria Donofrio, Felice Giangaspero, Daniel Picard, Marc Remke, Livia Garzia, Craig Daniels, Olivier Delattre, Fredrik J Swartling, William A Weiss, Francesco Salvatore, Roberto Fattorusso, Louis Chesler, Michael D Taylor, Giuseppe Cinalli, Massimo Zollo, Metastatic group 3 medulloblastoma is driven by PRUNE1 targeting NME1–TGF-β–OTX2–SNAIL via PTEN inhibition, Brain, Volume 141, Issue 5, May 2018, Pages 1300–1319, https://doi.org/10.1093/brain/awy039
Close - Share Icon Share
Abstract
Genetic modifications during development of paediatric groups 3 and 4 medulloblastoma are responsible for their highly metastatic properties and poor patient survival rates. PRUNE1 is highly expressed in metastatic medulloblastoma group 3, which is characterized by TGF-β signalling activation, c-MYC amplification, and OTX2 expression. We describe the process of activation of the PRUNE1 signalling pathway that includes its binding to NME1, TGF-β activation, OTX2 upregulation, SNAIL (SNAI1) upregulation, and PTEN inhibition. The newly identified small molecule pyrimido-pyrimidine derivative AA7.1 enhances PRUNE1 degradation, inhibits this activation network, and augments PTEN expression. Both AA7.1 and a competitive permeable peptide that impairs PRUNE1/NME1 complex formation, impair tumour growth and metastatic dissemination in orthotopic xenograft models with a metastatic medulloblastoma group 3 cell line (D425-Med cells). Using whole exome sequencing technology in metastatic medulloblastoma primary tumour cells, we also define 23 common ‘non-synonymous homozygous’ deleterious gene variants as part of the protein molecular network of relevance for metastatic processes. This PRUNE1/TGF-β/OTX2/PTEN axis, together with the medulloblastoma-driver mutations, is of relevance for future rational and targeted therapies for metastatic medulloblastoma group 3.
Introduction
Medulloblastoma is an embryonal tumour of the cerebellum that constitutes ∼20% of all primary CNS tumours in children under 14 years (Gajjar et al., 2004). Recent integrative genomics and mRNA expression profiling have demonstrated that medulloblastoma is not a single disease entity (Kool et al., 2008; Gibson et al., 2010; Cho et al., 2011; Northcott et al., 2011; Remke et al., 2011; Rausch et al., 2012). Subclassification of the intertumoral heterogeneity of medulloblastoma into 12 subtypes was defined through combination of gene expression and methylation data (Cavalli et al., 2017): two WNT (α, β); four SHH (α, β, γ, δ); three Group 3 (α, β, γ); and three Group 4 (α, β, γ). Medulloblastoma with activated WNT signalling (MBWNT) has favourable patient outcome and very low frequency of metastatic disease at diagnosis, while MBSHH, MBGroup3 and MBGroup4 are considered to have high metastatic potential.
Here we focus on MBGroup3. MBGroup3-α shows higher chromosome -8q loss (MYC locus: 8q24), as part of ‘high-risk’ MBGroup3. MBGroup3-β shows loss of chromosomes -1 and -9. The MBGroup3-γ subtype, with high metastatic potential and poor survival rates (independent of MYC amplification), has a trend for i17q enrichment, gain of chromosome -1q, and higher frequency of chromosome -8q gain (Cavalli et al., 2017).
The tendency for local invasion in MBGroup3 leads to wide dissemination through the CSF in the leptomeningeal space (40% of children at diagnosis, and most tumours at recurrence). Given the propensity of medulloblastoma to disseminate throughout the neuroaxis, standard multi-modality management includes maximal safe resection followed by craniospinal irradiation, with radiation boost to the involved field or posterior fossa, and chemotherapy (cisplatin/etoposide; vincristine/cyclophosphamide) for 3-year-old affected children (Tarbell et al., 2013; Massimino et al., 2016).
Unfortunately, the molecular events underlying this dissemination remain elusive for metastatic medulloblastoma. Mutations in genes and chromosomal aberrations, genomic instability and chromothripsis are also known to occur as early events in many cases of MBGroup3/4. New studies into genetics and treatment of metastatic medulloblastoma are in great demand (Zollo, 2013).
Here, we investigated PRUNE1, a member of the DHH (Asp-His-His) protein superfamily with an exopolyphosphatase activity for short-chain over long-chain inorganic polyphosphates (Tammenkoski et al., 2008). Its overexpression promotes cell motility, cancer progression and metastasis (Forus et al., 2001; D'Angelo et al., 2004). PRUNE1 is an unfolded multi-domain adaptor protein that can interact with binding partners and modulate signalling cascades, including WNT and TGF-β signalling, and has been shown to serve as nodal points in regulation of many cellular activities (Carotenuto et al., 2014). The WNT and TGF-β pathways are cell–cell signalling systems that control a plethora of processes, from embryonic development and cell proliferation, differentiation and migration, to tissue homeostasis, stem-cell behaviour, tissue regeneration, and cancers (Massague, 2008). TGF-β is also known to regulate systemic immune surveillance of the tumour host by controlling immune responses, and it maintains immune homeostasis through its impact on proliferation, differentiation and survival of multiple immune cell lineages (Pickup et al., 2013).
Studies using nuclear magnetic resonance have identified and structured the physical interactions of PRUNE1 with NME1 (Diana et al., 2013). PRUNE1 and NME1 are preferentially expressed during brain development (Carotenuto et al., 2006; Galasso and Zollo, 2009). Furthermore, mutations in PRUNE1 were recently identified in families with autosomal recessive inheritance affected by microcephaly and progressive encephalopathy–hypsarrhythmia–optical atrophy (PEHO) syndrome, with cerebellar atrophy showing gain-of-function PRUNE1 mutation effects on microtubule polymerization during mitosis in neural cells (Zollo et al., 2017).
Here, we identified a role for PRUNE1 (together with other functionally related genes) in metastatic MBGroup3 through minigene ontology analyses using several gene expression assays in a cohort of subjects with medulloblastoma. We demonstrate here that through binding to NME1, PRUNE1 enhances the canonical TGF-β signalling pathway mediated by SMAD2/3 in MBGroup3, which is known to be involved in tumour invasion and metastasis. Together with functional studies in immortalized and primary medulloblastoma cells, we propose a novel mechanism of action for PRUNE1 through upregulation of OTX2 and inhibition of tumour suppressor PTEN. This is confirmed by immunopathological analyses in our medulloblastoma collection.
A competitive permeable peptide (CPP) that impairs PRUNE1/NME1 complex formation (Carotenuto et al., 2015; Ferrucci et al., 2018), together with a novel ‘anti-PRUNE1’ compound (AA7.1), provide inhibition of tumour progression and metastatic dissemination in orthotopic xenograft models with a metastatic MBGroup3 cell line (D425-Med cells). We provide evidence for their potential future application in therapies against metastatic dissemination of MBGroup3. Through next-generation sequencing (NGS), 23 deleterious gene variants (non-synonymous variants, NSVs) common to metastatic primary MBgroup3/4 cells were identified. We indicate their genetic predisposition to regulate tumour progression, provide the ‘soil’ for the assembly of a pro-metastatic niche, and control the actions of immune cells. Taken together, these data provide the basis for future rational and targeted therapies to treat these highly metastatic paediatric medulloblastoma tumours.
Materials and methods
Cell proliferation and migration assays for cell index
Cell proliferation
Human primary MBGroup3 cells (pMB7), MBGroup4 cells (pMB6) or D425-Med (5 × 103) were plated into single wells of the xCELLigence® E-plate 16 (ACEA Biosciences, #05469830001). Then AA7.1 (100 µM) or phosphate-buffered saline (PBS, vehicle) were added, with the proliferation rate determined by measuring electrical impedance changes at the bottom of each well, at 2-min intervals over 48 h.
Cell migration
D425-Med cells were infected with AdV-sh-PRUNE1, AdV-CPP, AdV-Mock [multiplicity of infection (MOI), 100] or treated with 100 µM AA7.1, harvested, washed with PBS, resuspended to 7 × 104 cells/ml in Iscove’s modified Dulbecco’s medium without foetal bovine serum (FBS). Each cell suspension was used in a single well of an xCELLigence® CIM-plate 16 (ACEA Biosciences, #05665817001). Cell migration was driven by a 10% FBS gradient, with 0% FBS used as negative control. Measurements were taken at 2-min intervals, of impedance changes across the electrodes at the bottoms of the wells, over 12 h.
Luciferase reporter assays
HEK-293T cells were transfected with TransIT-LT1 Transfection Reagent (Mirus Bio LLC, #MIR2300), according to the manufacturer instructions. The following plasmids were used: pGL4.14-4XSBE-CMV-luc2/Hygro, pCS-3XFLAG-PRUNE1, pCMV5B-SMAD4-HA, pCDNA-SMAD3-FLAG, and pcDNA-HA-NME1. After 48 h transfection, HEK-293T cells were lysed with Passive Lysis Buffer (Promega, #E1910), and the luciferase activity (Luc) in the cell lysates was determined following the manufacturer instructions of Dual Luciferase Reporter Assays (Promega, #E1910). Light emission was quantified with EnSpire Multimode, and luciferase values were calculated.
Next-generation sequencing in metastatic primary medulloblastoma
The NGS data are available at BioSample NCBI accessions. Two primary cell types were used: SAMN04528823 (pMB6 cells) and SAMN04528824 (pMB7 cells). For cell handling methodologies see the Supplementary material. Here, 1774 ‘coding NSVs’ were found in common with tumour samples from metastatic primary MBGroup3/4 cells (pMB7 and, pMB6, respectively). Deleterious loss-of-function NSVs were identified by ‘sorting intolerant from tolerant’ (SIFT score, 0.00–0.05), ‘polymorphism phenotyping’ (PolyPhen; score, 0.9–1.0), and ClinVar archives database ‘not benign’ (http://www.ncbi.nlm.nih.gov/clinvar/). A protein interaction network was generated using the Search Tool for the Retrieval of Interacting Genes/Proteins (STRING) database (combined score, set to 0.4), together with NME1 and PTEN. Here, 23 NSVs were included in the gene network identified.
Animal procedures and treatments
The Animal Research Reporting of In Vivo Experiments (ARRIVE) Guidelines were followed. The orthotopic intracerebellar xenograft models used athymic Nude-Foxn1nu male mice (4 weeks old) according to the methodology described by Asadzadeh et al. (2017) (Supplementary material). AA7.1 was given intraperitoneally at 60 mg/kg every 2 days. After 28 days, the mice were sacrificed and their blood and organs were collected for further analysis.
AA7.1 toxicity
The toxicity of AA7.1 was determined with 4-week-old Balb/c mice grouped according to body weight. Escalating doses of AA7.1 (15, 30, 60 mg/kg/day) or vehicle (PBS) were administered intraperitoneally, daily for 1 week. At sacrifice, the blood and organs were collected and evaluated. Haematological and biochemical markers of hepatic and renal functions were evaluated. Data are expressed as means from three mice within each treatment group.
Ethical approval for human and mouse use
Informed consent for the Italian human tumour tissue bank was approved by Azienda Ospedaliera di Rilievo Nazionale ‘Santobono-Pausillipon’; (DL. 121, 29-4-2010; date 27/3/2013). For the mice studies: Ministero della Sanità 546/2015 PR released to the Director of Studies, Massimo Zollo, 19/6/2015, art. 31 D.lgs. 26/2104.
Statistical analyses
The data representative of three independent experiments were analysed using unpaired two-tailed t-tests (Student’s t-tests). For all figures, error bars represent standard deviation (SD) of the mean, and P < 0.05 was considered significant. The distribution of the event-free survival was calculated according to the Kaplan–Meier method, using Statistical Package for Social Sciences (IBM-SPSS, Version 7.0). The correlation analysis was carried out through R2 (Genomics Analysis and Visualisation Platform; http://r2.amc.nl). The protein densitometry for western blotting quantification used was carried out using the ImageJ software to calculate the relative and normalized densities of peaks corresponding to the bands for proteins of interest, and those relative to the loading-control bands. Cell fluorescence was measured using ImageJ software, and the corrected total cell fluorescence (CTCF) was estimated according to the following formula, as described by McCloy et al. (2014): CTCF = integrated density − (area of selected cell × mean background fluorescence).
Results
PRUNE1 expression as a metastatic marker in medulloblastoma
PRUNE1 is known to initiate cell migration and metastasis (D'Angelo et al., 2004; Zollo et al., 2005; Oue et al., 2007; Noguchi et al., 2009). To investigate its role in metastatic medulloblastoma, gene expression data from public medulloblastoma datasets (Delattre, Gilbertson, Kool and Roth) were analysed using R2 Genomics Analysis and Visualisation Platform (http://r2.amc.nl). The data showed higher PRUNE1 expression in medulloblastoma samples compared to normal cerebellum (P = 1.6 × 10−9; Supplementary Fig. 1A). Among medulloblastoma molecular subgroups, PRUNE1 transcriptional levels were significantly higher in MBGroups3/4 compared to MBWNT and MBSHH, in two non-overlapping public datasets (Boston, P < 0.05; Cavalli, P < 0.001; Fig. 1A and B).
PRUNE1 expression in medulloblastoma. (A and B) RNA log2 expression of PRUNE1 derived from transcriptome analysis from the primary cohort of medulloblastoma (A; Boston cohort; n = 187; P < 0.05) and from the publicly available dataset of medulloblastoma (B; Cavalli; n = 763; P < 0.001), grouped according to the molecular and subtype disease variants. PRUNE1 was highly expressed in MBGroup3 and MBGroup4. Normal CB = normal cerebellum. (C) RNA log2 expression of PRUNE1 across medulloblastoma primary tumours [i.e. M1 (n = 176) versus M0 (n = 397)] in the publicly available Cavalli dataset (n = 573 tumours). There were higher PRUNE1 expression levels in metastatic M1 compared to M0 samples (P = 0.05). (D) Real-time PCR analysis on a ‘non-overlapping’ cohort of our medulloblastoma tissues from Italy, according to metastatic status at diagnosis (M0, n = 26; M1, n = 8). PRUNE1 expression levels were higher in metastatic M1 tumours compared to M0 (P = 0.05). See Supplementary Table 1 for real-time PCR expression data. (E) Kaplan-Meyer analysis for event-free survival (EFS) of MBGroup3 and MBGroup4 patients according to PRUNE1 expression levels. Patients who showed higher levels of PRUNE1 expression (n = 6) showed significantly shorter event-free survival compared to those with lower levels of PRUNE1 expression (n = 4) (P < 0.05). See Supplementary Table 1 for real-time PCR expression data. (F) RNA log2 expression of NME1 across the primary medulloblastoma samples from the publically available Cavalli dataset (n = 763; P < 0.001) grouped according to molecular subtypes. NME1 was highly expressed in MBGroup3-γ subtype.
Knowing that metastatic events are more common in MBGroup3 (46.5%) and MBGroup4 (29.7%) than in MBWNT (17.9%) and MBSHH (19.1%) (Cavalli et al., 2017), we analysed PRUNE1 expression levels by comparing metastatic and non-metastatic samples (i.e. M1 versus M0). There was significantly higher PRUNE1 expression in metastatic M1 tumours (n = 176) compared to M0 tumours (n = 397) (P = 0.05; Fig. 1C). Then our collection of primary medulloblastoma samples was used (44 samples; Table 1). Here, MBGroup3 (n = 7) negatively correlated with event-free survival (Kaplan–Meyer analysis, P = 0.04; Supplementary Fig. 1B). Real-time PCR showed PRUNE1 mRNA levels to be significantly higher in M1 tumours (n = 8) compared to M0 tumours (n = 26; P = 0.05; Fig. 1D and Supplementary Table 1). Strikingly, when MBGroup3/4 patients were stratified at diagnosis according to mRNA levels, those with higher PRUNE1 (n = 6) expression had significantly shorter event-free survival than those with lower PRUNE1 (n = 4) expression (P = 0.05; Fig. 1E and Supplementary Table 1, see also Supplementary material). Altogether, these data indicated that relapsing/metastatic medulloblastoma shows correlations with high levels of PRUNE1 in MBGroup3/4 primary tumours.
Clinical features of the patient included, as of September 2017
| Characteristic . | Measure . | Total population n (%)/datum . | According to risk group n (%)/datum . | |
|---|---|---|---|---|
| Standard . | High . | |||
| Total patients | 44 (100) | 18 (40.9) | 26 (59.1) | |
| Age at diagnosis (years) | Median | 6.00 | 7.29 | 3.41 |
| Range | 0.42–15.33 | 3.58–15.33 | 0.42–12.00 | |
| Gender | Male | 19 (43.19) | 6 (33.3) | 13 (50.0) |
| Female | 25 (56.81) | 12 (66.7) | 13 (50.0) | |
| Male/female ratio | 0.76 | 0.50 | 1.00 | |
| Metastatic status | M0 | 26 (59.09) | 18 (100) | 8 (30.8) |
| M1-M4 | 18 (40.9) | 0 (0.0) | 18 (69.2) | |
| Histopathological variant | Classic | 21 (47.72) | 12 (66.7) | 9 (34.6) |
| Desmoplastic/nodular | 18 (40.9) | 5 (27.8) | 13 (50.0) | |
| Large cell/anaplastic | 5 (11.37) | 1 (5.6) | 4 (15.4) | |
| Follow-up (September 2017) | Total | 38 (100) | 14 (36.8) | 24 (63.2) |
| Status | Alive, no evidence of disease | 22 (57.8) | 8 (57.1) | 14 (58.3) |
| Alive, with disease | 2 (5.26) | 0 (0) | 2 (8.3) | |
| Dead of disease | 14 (36.84) | 6 (42.9) | 8 (33.3) | |
| Characteristic . | Measure . | Total population n (%)/datum . | According to risk group n (%)/datum . | |
|---|---|---|---|---|
| Standard . | High . | |||
| Total patients | 44 (100) | 18 (40.9) | 26 (59.1) | |
| Age at diagnosis (years) | Median | 6.00 | 7.29 | 3.41 |
| Range | 0.42–15.33 | 3.58–15.33 | 0.42–12.00 | |
| Gender | Male | 19 (43.19) | 6 (33.3) | 13 (50.0) |
| Female | 25 (56.81) | 12 (66.7) | 13 (50.0) | |
| Male/female ratio | 0.76 | 0.50 | 1.00 | |
| Metastatic status | M0 | 26 (59.09) | 18 (100) | 8 (30.8) |
| M1-M4 | 18 (40.9) | 0 (0.0) | 18 (69.2) | |
| Histopathological variant | Classic | 21 (47.72) | 12 (66.7) | 9 (34.6) |
| Desmoplastic/nodular | 18 (40.9) | 5 (27.8) | 13 (50.0) | |
| Large cell/anaplastic | 5 (11.37) | 1 (5.6) | 4 (15.4) | |
| Follow-up (September 2017) | Total | 38 (100) | 14 (36.8) | 24 (63.2) |
| Status | Alive, no evidence of disease | 22 (57.8) | 8 (57.1) | 14 (58.3) |
| Alive, with disease | 2 (5.26) | 0 (0) | 2 (8.3) | |
| Dead of disease | 14 (36.84) | 6 (42.9) | 8 (33.3) | |
The tumour samples were from the Italian tissue cohort bank of patients enrolled at Santobono Hospital in Naples between 2006 and 2013. Patients were coded as ‘standard risk’ or ‘high risk’ as described in the Supplementary material. Among 44 tumour samples, because of tissue degradation or poor RNA quality, 36 were used for real-time PCR to analyse PRUNE1 expression (data related to Fig. 1D and E), and 26 were used for molecular subgrouping and classification (data related to Supplementary Fig. 1B).
Clinical features of the patient included, as of September 2017
| Characteristic . | Measure . | Total population n (%)/datum . | According to risk group n (%)/datum . | |
|---|---|---|---|---|
| Standard . | High . | |||
| Total patients | 44 (100) | 18 (40.9) | 26 (59.1) | |
| Age at diagnosis (years) | Median | 6.00 | 7.29 | 3.41 |
| Range | 0.42–15.33 | 3.58–15.33 | 0.42–12.00 | |
| Gender | Male | 19 (43.19) | 6 (33.3) | 13 (50.0) |
| Female | 25 (56.81) | 12 (66.7) | 13 (50.0) | |
| Male/female ratio | 0.76 | 0.50 | 1.00 | |
| Metastatic status | M0 | 26 (59.09) | 18 (100) | 8 (30.8) |
| M1-M4 | 18 (40.9) | 0 (0.0) | 18 (69.2) | |
| Histopathological variant | Classic | 21 (47.72) | 12 (66.7) | 9 (34.6) |
| Desmoplastic/nodular | 18 (40.9) | 5 (27.8) | 13 (50.0) | |
| Large cell/anaplastic | 5 (11.37) | 1 (5.6) | 4 (15.4) | |
| Follow-up (September 2017) | Total | 38 (100) | 14 (36.8) | 24 (63.2) |
| Status | Alive, no evidence of disease | 22 (57.8) | 8 (57.1) | 14 (58.3) |
| Alive, with disease | 2 (5.26) | 0 (0) | 2 (8.3) | |
| Dead of disease | 14 (36.84) | 6 (42.9) | 8 (33.3) | |
| Characteristic . | Measure . | Total population n (%)/datum . | According to risk group n (%)/datum . | |
|---|---|---|---|---|
| Standard . | High . | |||
| Total patients | 44 (100) | 18 (40.9) | 26 (59.1) | |
| Age at diagnosis (years) | Median | 6.00 | 7.29 | 3.41 |
| Range | 0.42–15.33 | 3.58–15.33 | 0.42–12.00 | |
| Gender | Male | 19 (43.19) | 6 (33.3) | 13 (50.0) |
| Female | 25 (56.81) | 12 (66.7) | 13 (50.0) | |
| Male/female ratio | 0.76 | 0.50 | 1.00 | |
| Metastatic status | M0 | 26 (59.09) | 18 (100) | 8 (30.8) |
| M1-M4 | 18 (40.9) | 0 (0.0) | 18 (69.2) | |
| Histopathological variant | Classic | 21 (47.72) | 12 (66.7) | 9 (34.6) |
| Desmoplastic/nodular | 18 (40.9) | 5 (27.8) | 13 (50.0) | |
| Large cell/anaplastic | 5 (11.37) | 1 (5.6) | 4 (15.4) | |
| Follow-up (September 2017) | Total | 38 (100) | 14 (36.8) | 24 (63.2) |
| Status | Alive, no evidence of disease | 22 (57.8) | 8 (57.1) | 14 (58.3) |
| Alive, with disease | 2 (5.26) | 0 (0) | 2 (8.3) | |
| Dead of disease | 14 (36.84) | 6 (42.9) | 8 (33.3) | |
The tumour samples were from the Italian tissue cohort bank of patients enrolled at Santobono Hospital in Naples between 2006 and 2013. Patients were coded as ‘standard risk’ or ‘high risk’ as described in the Supplementary material. Among 44 tumour samples, because of tissue degradation or poor RNA quality, 36 were used for real-time PCR to analyse PRUNE1 expression (data related to Fig. 1D and E), and 26 were used for molecular subgrouping and classification (data related to Supplementary Fig. 1B).
The PRUNE1/NME1 protein complex is involved in metastatic dissemination (D'Angelo et al., 2004; Carotenuto et al., 2013). To determine whether this complex can affect medulloblastoma metastatic progression, we investigated expression of NME1 in medulloblastoma samples using the Cavalli dataset. There was overexpression of NME1 in MBGroup3-γ (Cavalli, P < 0.001; Fig. 1F). In addition, NME1/NME2 were more expressed in metastatic samples (i.e. M1 versus M0) (Cavalli, P = 6.9 × 10−03; Supplementary Fig. 1C).
PRUNE1 and NME1 protein levels were also investigated among different medulloblastoma cell lines using western blotting. These data showed higher levels of PRUNE1 and NME1, including activated phospho-Ser120-122-125-NME1 (Garzia et al., 2008), whereby serine phosphorylation mediates the protein-protein complex, in all the MBGroup3 cells (D425-Med, D341-Med, D283-Med cells) compared to MBSHH (UW-288, DAOY, ONS-76 cells) (Supplementary Fig. 1D and E).
These findings were confirmed by significant positive correlation between PRUNE1 and NME1/NME2 in the subset of metastatic patients (Cavalli: r = 0.23, P = 2.1 × 10−03; Supplementary Fig. 1F). These data showed that PRUNE1 and its protein partner NME1 are overexpressed in metastatic MBGroup3, which suggested that these genes/proteins might influence those metastatic molecular processes.
PRUNE1 and OTX2 expression correlate with canonical TGF-β signalling
Gene expression data followed by mini-gene-ontology analysis were used to identify other genes correlated to PRUNE1 in metastatic MBGroup3 (Supplementary material). This approach identified the OTX2, CYFIP1 and GLI2 genes (involved in neurogenesis and neuronal differentiation processes), with OTX2 as the most relevant (Supplementary Table 2). OTX2 is highly expressed in MBGroup3 (Boulay et al., 2017), and it can regulate cell proliferation, growth, self-renewal, migration and survival processes (Adamson et al., 2010; Bai et al., 2012; Bunt et al., 2013). PRUNE1 and OTX2 were positively correlated in primary medulloblastoma samples (Cavalli: r = 0.455, P = 3.2 × 10−40; Supplementary Fig. 2A).
OTX2 is known to act in canonical TGF-β signalling in zebrafish (through SMAD2/3) (Jia et al., 2009). Similarly, TGF-β1 cytokine (TGFB1) and its serine/threonine kinase receptor type 1 (TGFBR1) also positively correlated to OTX2 expression (Cavalli: r = 0.361, P = 6.3 × 10−25; r = 0.286, P = 8.5 × 10−16, respectively; Supplementary Fig. 2B and C). Indeed, TGFB1 and its receptor TGFBR1 were mainly expressed in medulloblastoma samples compared to normal cerebellum, and were overexpressed in metastatic (M1) MBGroup3 (Cavalli: P = 3.0 × 10−4; P = 0.003, respectively; Supplementary Fig. 2D).
The downstream effectors of canonical TGF-β signalling (i.e. SMAD2, SMAD3 and SMAD4), are known to act at the interplay between oncogenic and tumour-suppressor actions during tumour progression (Massague, 2008). With similar searches within the MBGroup3 tumours, both R-SMADs (SMAD2, SMAD3) and co-SMAD (SMAD4) were highly expressed in the most aggressive and metastatic MBGroup3-α and MBGroup3-γ subtypes (Cavalli: SMAD2, P = 2.1 × 10−5; SMAD3, P = 2 × 10−5; SMAD4, P = 6.7 × 10−3; Supplementary Fig. 2E). These provided sufficient evidence that canonical TGF-β signalling is involved in the metastatic behaviour of MBGroup3 tumours.
As MYC gene family members (e.g. c-MYC, N-MYC) are known to be significant contributors to initiation, maintenance and progression of MBGroup3 (Cavalli et al., 2017), we investigated potential correlations across PRUNE1, c-MYC, N-MYC and TGF-β. In the Cavalli dataset, c-MYC was overexpressed in MBGroup3 and MBWNT subtypes (P = 1.1 × 10−158; Supplementary Fig. 3A), although not significantly correlated to metastatic status (M1) of medulloblastoma patients (M0: n = 397; M1: n = 176; data not shown). In contrast, overexpression of N-MYC was found in MBSHH and MBWNT (P = 3.5 × 10−132; Supplementary Fig. 3B) and in non-metastatic patients (M0; P = 5.7 × 10−04; Supplementary Fig. 3C). Furthermore, c-MYC positively correlated (as ‘weak’) to both PRUNE1 (r = 0.173, P = 1.5 × 10−06) and OTX2 (r = 0.255, P = 18.0 × 10−13), and showed ‘moderate’ positive association with TGFB1 (r = 0.420, P = 5.8 × 10−34; Supplementary Table 3). In contrast, N-MYC negatively correlated to PRUNE1 (r = −0.439, P = 3.3 × 10−37), TGFB1 (r = −0.258, P = 4.6 × 10−13) and OTX2 (r = −0.556, P = 3.1 × 10−63; Supplementary Table 3). Overall, c-MYC expression correlated to MBGroup3 and MBWNT, while N-MYC expression correlated to MBSHH and MBWNT, and mainly to M0 status.
We then evaluated whether OTX2 is a downstream gene target of canonical TGF-β signalling within the PRUNE1/NME1 protein complex in metastatic MBGroup3, as this axis is independent of c-MYC amplification status (Cavalli et al., 2017). To this end, taking into account the gene-expression data (R2; http://hgserver1.amc.nl/; defined as the ‘Kool’ dataset), D283-Med cells were selected as the best cell model to represent MBGroup3 in this study of the mechanism of action of PRUNE1/NME1, compared to the other available MBgroup3 cell lines (Fig. 2A). These D283-Med cells were defined as having loss of copy number variations within the genomic region of the PRUNE1, OTX2, c-MYC, N-MYC and TP53 genes, and lower expression of the NME1 and NME2 genes compared to the other MBgroup3 cell lines for TP53 status (Ivanov et al., 2016).
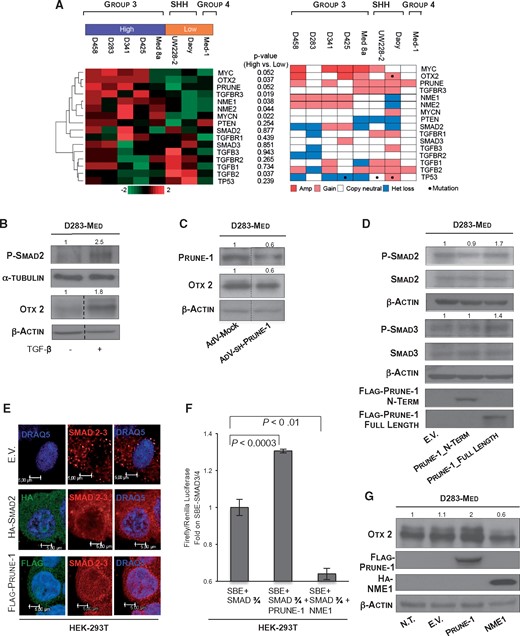
Role of PRUNE1 in the canonical TGF-β signalling cascade and OTX2 upregulation. (A) Medulloblastoma cell lines were grouped and cross-referenced with c-MYC expression levels and with copy number variations (e.g. amplification and/or gain; as from the data from the COSMIC and cBioPortal datasets). Left: Transcriptome analysis of immortalized medulloblastoma cell lines used to characterize PRUNE1-associated genes (R2: http://hgserver1.amc.nl/). Cell lines in the ‘high group’ (purple highlight) were characterized either by c-MYC amplification or by high expression of c-MYC, whereas the remaining cell lines were in the ‘low group’ (orange highlight). Hierarchical clustering was performed on the genes indicated and categorized according to ‘high versus low’ groups. Right: Copy number data showing status of the genes indicated (red, amplified; pink, gained; white, copy neutral; blue, heterozygous loss). Mutational status is indicated by the dots. Sources for data: published TP53 status for all samples (Ivanov et al., 2016); complete NGS-WES data available for MBGroup3 D341-Med and MBSHH DAOY (http://www.cbioportal.org), MBGroup3 D283-Med (http://cancer.sanger.ac.uk/cosmic). The peritoneal metastatic D283-Med cell line was selected as the representative MBGroup3 model for this study, because of lack of copy number variations (e.g. amplifications or gains, loss of heterozygosity, mutations) in the genomic region of the PRUNE1, OTX2, TP53, c-MYC and N-MYC genes, and lower relative expression levels of the NME1 and NME2 genes compared to the other MBgroup3 cell lines. (B) Representative western blot showing upregulation of phospho-Ser467-SMAD2 (2.5-fold) and OTX2 (1.8-fold) protein levels in cell lysates from MBGroup3 D283-Med cells treated with TGF-β1 cytokine. α-Tubulin and β-actin levels were used as loading controls. (C) Representative western blot of cell lysates of D283-Med cells under PRUNE1 silencing using a short hairpin ‘sh-PRUNE1’ carried by adenoviral particles (AdV-sh-PRUNE1), with D283-Med cells infected with Adv-Mock as negative controls. Both PRUNE1 and OTX2 protein levels were decreased in D283-Med cells infected with AdV-sh-PRUNE1, in comparison to those infected with AdV-Mock (PRUNE1, 0.6-fold; OTX2, 0.6-fold). β-Actin levels were used as the loading control. (D) Representative western blot on cell lysates from MBGroup3 D283-Med cells transiently transfected with plasmids containing FLAG-tagged amino-terminus-PRUNE1 (Prune-1_N-Term), FLAG-tagged full-length PRUNE1 (Prune-1_Full-Length), and empty vector (E.V.) as negative control. The phosphorylated-Ser467-SMAD2 (1.7-fold) and phosphorylated-Ser423/425-SMAD3 (1.4-fold) protein levels increased in cells overexpressing full-length PRUNE1 compared to those transfected with Prune-1_N-Term and empty vector (E.V.) plasmids. β-Actin levels were used as loading control. (E) Confocal immunofluorescence analysis showing subcellular distribution of R-SMADs (i.e. SMAD2/3; red) in HEK-293T cells transiently transfected with full-length PRUNE1 (green), SMAD2 (green), and empty vector (E.V.) plasmids (positive and negative controls, respectively). Similar nuclear staining for SMAD2/3 complex (red) was shown in HEK-293T cells overexpressing PRUNE1 and SMAD2, in comparison to empty vector control plasmid, which showed cytoplasmic distribution for R-SMADs. Nuclei were stained with DRAQ5 (as blue staining). Scale bars = 5 µm. Assays were performed as independent triplicates, with photographs taken as representative of all of these experiments. (F) Transactivation assays in transiently transfected HEK-293T cells using firefly luciferase constructs that contained Smad binding elements (SBE), together with plasmids containing SMAD3 and SMAD4 (as positive controls) in combination with full-length PRUNE1 or NME1, as indicated. Firefly Luciferase (Luc) activity is shown as fold on positive control. PRUNE1 enhanced the TGF-β signalling cascade, as an increase in Luc activity (1.3-fold; P < 0.0003). In contrast, NME1 inhibited TGF-β signalling activation, as seen here (0.6-fold; P < 0.01). (G) Representative western blot showing upregulation and downregulation of OTX2 levels in D283-Med cells transiently transfected with full-length PRUNE1 and HA-tagged-NME1, respectively, in comparison to D283-Med cells transfected with empty vector (E.V.) plasmid and non-transfected (N.T.) cells. The densitometric analyses showed up-regulation of OTX2 (2-fold) in D283-Med cells expressing PRUNE1, and down-regulation (0.6-fold) in those overexpressing NME1. β-Actin levels were used as the loading control.
Western blotting was also used for evaluation of OTX2 expression in several medulloblastoma cell lines, whereby OTX2 levels were higher in MBGroup3 cells compared to MBSHH cell lines (Supplementary Fig. 3D), thus confirming the gene expression data.
To define the modulation of OTX2 levels mediated by TGF-β, D283-Med cells were treated with this cytokine (5 ng/ml) or vehicle, as negative control. TGF-β promoted increased activated SMAD2 (phosphorylated at Ser467) and upregulated OTX2 expression (Fig. 2B). In the control, SB525334 (inhibitor of TGF-β receptor type 1) decreased OTX2 protein levels and phosphorylated SMAD2 (data not shown). To determine whether these increased OTX2 levels were due to activation of the canonical TGF-β/SMAD2-mediated pathway, transient SMAD2 overexpression was performed in D283-Med cells (Supplementary Fig. 3E). Taken together, these data showed that OTX2 is regulated by TGF-β via the TGF-β/SMAD2 canonical pathway.
We then investigated whether PRUNE1 influences this TGF-β-induced OTX2 axis. PRUNE1 expression was downregulated in D283-Med cells to determine how this axis behaved with low PRUNE1 expression. Interestingly, this resulted in downregulation of OTX2 (Fig. 2C). These findings further suggested that activation of TGF-β positively correlated to OTX2 expression, with PRUNE1 silencing resulting in inhibition of OTX2 expression. We therefore hypothesized that PRUNE1 modulates OTX2 expression via the canonical TGF-β pathway. The activation levels of SMAD2 and SMAD3 were thus analysed in terms of their phosphorylation status and subcellular distribution upon PRUNE1 overexpression. Phosphorylated-Ser467-SMAD2 and phosphorylated-Ser423/425-SMAD3 protein levels increased with full-length PRUNE1 in transfected D283-Med cells (Fig. 2D). These findings suggested that this regulation occurs at the carboxyl-terminus region of PRUNE1. Then immunofluorescence microscopy was used to show that transient overexpression of PRUNE1 increased the levels of the nuclear SMAD2/3 protein complex in HEK-293T cells, thus inducing their nuclear accumulation (positive control: SMAD2-transfected cells; Fig. 2E). Overall, these data supported the concept that PRUNE1 participates in the canonical TGF-β signalling cascade through nuclearization of the SMAD2/3 protein complex, as an action mediated by the carboxyl-terminal region of PRUNE1, as the region of interaction with NME1 (Diana et al., 2013).
We investigated further the role of PRUNE1 on transcriptional activation through the SMAD binding elements (SBEs; Morikawa et al., 2013) using HEK-293T cells, because of their negligible levels of endogenous PRUNE1 expression (Carotenuto et al., 2014). These data indicated that PRUNE1 enhances SMAD-transactivation induced by SMAD3 and SMAD4 (as positive controls; 1.3-fold; P < 0.0003; Fig. 2F and Supplementary Fig. 3F). In summary here, we have defined a novel role for PRUNE1 as a promoter of the SMAD2/3-mediated canonical TGF-β pathway.
Of note, NME1 has been reported to negatively regulate TGF-β signalling through a physical association with serine/threonine kinase receptor associated protein (STRAP), which is known to prevent activation of SMAD effector proteins (Zhao et al., 2013). To gain insight into the mechanisms of PRUNE1/TGF-β/OTX2 axis regulation, we investigated the effects of NME1 on negative regulation of TGF-β signalling in medulloblastoma. Here, NME1 overexpression impaired the canonical TGF-β pathway, with reduction of SMAD transactivation (0.64-fold; P < 0.01; Fig. 2F and Supplementary Fig. 3F) in HEK-293T cells. Furthermore, this overexpression resulted in decreased levels of phosphorylated-Ser467-SMAD2 (Supplementary Fig. 3G) and OTX2 (Fig. 2G), which thus confirmed the inhibitory effects of NME1 on the TGF-β canonical pathway also in a tumorigenic MBGroup3 cell line. In contrast, the overexpression of STRAP did not affect OTX2 expression levels (Supplementary Fig. 3H). This further suggested that through its binding to NME1, PRUNE1 impairs NME1 inhibitory activity on the TGF-β canonical pathway, thus enhancing TGF-β signalling. Thus, this PRUNE1 action did not occur via STRAP-dependent mechanisms in these D283-Med cells.
PRUNE1 sustains oncogenic pathways in medulloblastoma through PTEN repression
PRUNE1 has been shown to be expressed in liver metastases generated from colorectal tumours induced by epithelial–mesenchymal transition (EMT) (Hashimoto et al., 2016). EMT also induces chemo-resistance in medulloblastoma (Asuthkar et al., 2011). For these reasons, we investigated whether PRUNE1-induced TGF-β signalling activates EMT in D283-Med cells (using modulation of E-cadherin and N-cadherin as EMT markers). Here, the overexpression of PRUNE1 decreased E-cadherin levels, while N-cadherin levels did not change (negative control: empty vector; Fig. 3A). Also, PRUNE1 activated nuclear TGF-β (i.e. phosphorylated-Ser467-SMAD2) and EMT markers (i.e. N-cadherin, E-cadherin) at the invasive front of MBGroup3 tumour tissues (Fig. 3B). Histological analysis showed high expression of both PRUNE1 and phosphorylated/nuclear SMAD2. Most importantly, there was strong N-cadherin-positive staining with very low but detectable levels of E-cadherin (Fig. 3B), thus further confirming the activation of EMT. Further histological analysis showed that all MBGroup3 samples from our cohort were positive for N-cadherin staining at the invasive front (11/11 samples), with lower expression of E-cadherin (10/11 samples; Fig. 3B and Supplementary Fig. 3I). Taken together, PRUNE1 contributes to EMT and metastatic dissemination in MBGroup3.
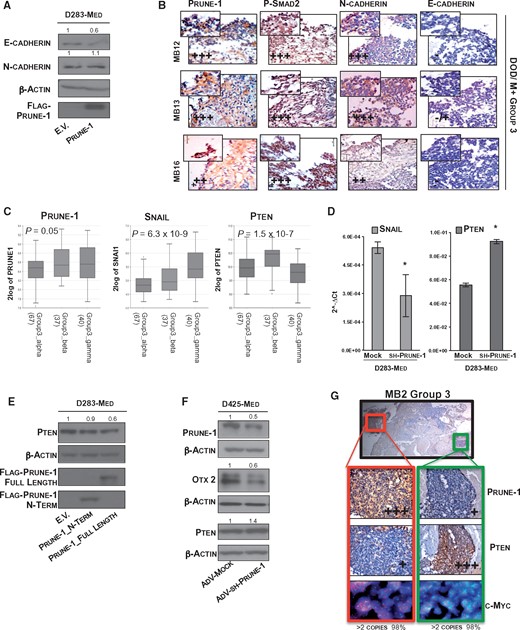
Activation of the AKT signalling pathway through PTEN inhibition mediated by PRUNE1. (A) Representative western blot with antibodies against epithelial-mesenchymal transition (EMT) markers (i.e. E-cadherin, N-cadherin) of cell lysates from D283-Med cells (using only adherent cells) after 48 h transfection with empty vector (E.V.) and full-length PRUNE1. β-Actin levels were used as the loading control. Densitometric measurements indicated decreased E-cadherin levels (0.6-fold) in PRUNE1-expressing cells compared to empty vector control, while N-Cadherin levels were unchanged here. (B) Immunohistochemistry using sequential sections of primary tumour tissue from MBGroup3 patients in our cohort with poor prognosis (M+ = metastatic MB; DOD = dead of disease), using anti-PRUNE1, anti-phospho-Ser467-SMAD2, anti-N-cadherin and anti-E-cadherin antibodies. Histopathological grading system (+++ = high grade; ++ = intermediate grade; + = low grade) showed levels of expression and distribution of the markers within the tumour sections. Nuclear phospho-SMAD2 and cytoplasmic PRUNE1 staining are shown with similar levels of expression. In tissue sections, nuclear staining of phosphorylated-SMAD2, together with positive immunoreactivity to N-cadherin and negativity to E-cadherin were seen next to the border at the tumour front (×40 magnification). (C) RNA log2 expression of PRUNE1, SNAIL (SNAI1) and PTEN across primary MBGroup3 tumour samples grouped according to molecular subtypes (i.e. α, β, γ) in the publicly available dataset (Cavalli, n = 144). Higher expression levels of PRUNE1 and SNAIL (SNAI1), and lower expression of PTEN, were seen in metastatic MBGroup3-γ. (D) Real-time PCR analysis showing expression levels of SNAIL (SNAI1) and PTEN in MBGroup3 D283-Med cells infected with adenovirus particles carrying sh-RNA for PRUNE1 (AdV-sh-PRUNE1), in comparison to AdV-Mock as control. Statistically significant reduction in SNAIL and increase in PTEN mRNA expression levels were seen in D283-Med cells upon PRUNE1 silencing, compared to cells infected with AdV-Mock (P < 0.05). (E) Representative western blot of cell lysates from MBGroup3 D283-Med cells transiently transfected with plasmids containing amino-terminus PRUNE1 (Prune-1_N-Term), full-length PRUNE1 (Prune-1_Full-Length) and with empty vector as control. Decreased PTEN protein levels (0.6-fold) were seen in D283-Med cells transiently overexpressing full-length PRUNE1, compared to those transfected with Prune-1_N-Term empty vector plasmids. β-Actin levels were used as loading control. (F) Representative western blot with indicated antibodies of protein lysates from MBGroup3 D425-Med cells infected with adenovirus carrying particles with AdV-sh-PRUNE1 or empty vector (Adv-Mock) as control. Data show downregulation of PRUNE1 (0.5-fold) and OTX2 (0.6-fold) and upregulation of PTEN (1.4-fold) in AdV-sh-PRUNE1 infected cells compared to AdV-Mock. (G) Metastatic MBgroup3 tumour (sample MB2), which showed an inverse correlation of positive staining (i.e. high PRUNE1, low PTEN levels; and vice versa) in other tissue regions of the cerebellar tumour. c-MYC (red) gene copy number is also shown (>2 copies).
SNAIL (encoded by SNAI1) has a key role in EMT, in both development and tumour progression (Lamouille et al., 2014). Moreover, OTX2 was reported to be tightly interconnected with SNAIL in the Ciona intestinalis embryo animal model (Kubo et al., 2010). With direct in vitro effects of PRUNE1 on TGF-β ruled out, we further investigated the impact of PRUNE1 on TGF-β signalling with the hypothesis that PRUNE1 contributes to metastatic dissemination by promoting EMT through SNAIL expression (Batlle et al., 2013). Interestingly, higher expression levels of PRUNE1 and SNAI1 were seen in the MBGroup3-γ subtype (PRUNE1, P = 0.05; SNAIL, P = 6.3 × 10−9; Fig. 3C), which suggested a functional relationship in these tumours. Furthermore, SNAIL has been reported to inhibit PTEN transcription (Escriva et al., 2008), to regulate neuronal progenitor cells in cerebellum (Zhu et al., 2017), and to have a ‘gate-keeper’ function in a murine model of MBSHH (Metcalfe et al., 2013). Of importance, lower PTEN expression was seen in the metastatic MBGroup3-γ subtype (P = 1.5 × 10−7; Fig. 3C).
For the above reasons, we investigated how PRUNE1 might negatively regulate PTEN expression via SNAIL. Here, PRUNE1 silencing significantly impaired SNAIL and upregulated PTEN mRNA expression in D283-Med cells (Fig. 3D). Remarkably, the overexpression of full-length PRUNE1 in D283-Med cells decreased PTEN levels (Fig. 3E). To confirm the inverse correlation between PRUNE1 and PTEN, AKT signalling was also investigated. Consistent with the previous data, PRUNE1-overexpressing D283-Med cells increased the phosphorylation of phospho-Ser473-AKT (Supplementary Fig. 3L). Together, these data indicated that PRUNE1 can act through the canonical TGF-β signalling cascade to induce SNAIL, repress PTEN expression and activate phosphatidylinositol-4,5-bisphosphate 3-kinase (PI3K) cascade.
We further validated this axis in another MBGroup3 cell model that carried TP53 mutation and c-MYC amplification (D425-Med cells). As expected, silencing PRUNE1 promoted reduction of OTX2 and upregulation of PTEN (Fig. 3F).
The inverse correlation between PRUNE1 and PTEN was further evaluated by immunohistochemistry in sequential sections from paraffin-embedded tumour samples from medulloblastoma patients representative of the four molecular groups. This confirmed high expression of PRUNE1 with lower PTEN levels in metastatic MBGroup3 (pMB7 and pMB9) and MBGroup4 (pMB6 and pMB8) compared to MBSHH and MBWNT (pMB3 and pMB4, respectively) (Supplementary Fig. 4A). The same inverse correlation between PRUNE1 and PTEN was observed in other MBGroup3 samples from our tissue cohort (Supplementary Fig. 4B). Interestingly, within the same tumour tissue, extreme marker heterogeneity was detected throughout, with the consistent finding of higher PRUNE1 expression with lower PTEN staining (Fig. 3G). We also looked for chromosomal abnormalities with a specific c-MYC probe, which showed that there were no significant copy-number variations in these tissue sections (Fig. 3G, Supplementary Fig. 4B and Supplementary material). Taken together, these data show extreme heterogeneity in marker expression within the same MBGroup3 tumours.
Considering our tumour samples, and using pathological scoring analyses (Louis et al., 2014), a high percentage of metastatic (M+) MBGroup3 tumours (n = 12) showed inverse expression between PRUNE1 and PTEN (i.e. PRUNE1High/PTENlow; n = 7/12; Supplementary Fig. 4A and B).
In conclusion here, PRUNE1 overexpression enhances TGF-β signalling via SMAD2/3 and OTX2, which results in PI3K pathway activation through SNAIL-mediated PTEN repression, thus enhancing EMT and metastases in MBGroup3. These findings prompted us to develop an in vivo model as a control within the interaction asset of PRUNE1/NME1, to then enhance the TGF-β canonical pathway.
A cell CPP designed to block PRUNE1/NME1 interactions inhibits medulloblastoma metastatic progression
The PRUNE1/NME1 interaction is known to be mediated by casein kinase 1-mediated phosphorylation of NME1 on its Ser120-122-125 residues (Garzia et al., 2008; Ferrucci et al., 2018). Here, we initially demonstrated that the carboxyl-terminus domain of PRUNE1 is sufficient to bind NME1 also in MBGroup3 (i.e. D283-Med cells) in an immunoprecipitation assay (Supplementary Fig. 4C). Then, the cells treated with a selective cell-permeant ATP-competitive inhibitor of casein kinase 1 (D4476), resulted in inhibition of OTX2 expression (Supplementary Fig. 4D).
We then investigated the actions of a competitive permeable peptide (CPP) in D283-Med and D425-Med cells using the adenoviral strategy described by Carotenuto et al. (2015). Reduced expression of phosphorylated-Ser(120-122-125)-NME1 and downregulation of phosphorylated-SMAD2 and OTX2 were observed (Fig. 4A). There was also decreased migration rate of D425-Med cells (Fig. 4B), which indicated the importance of the inhibition of the pro-migratory phenotype mediated by CPP.
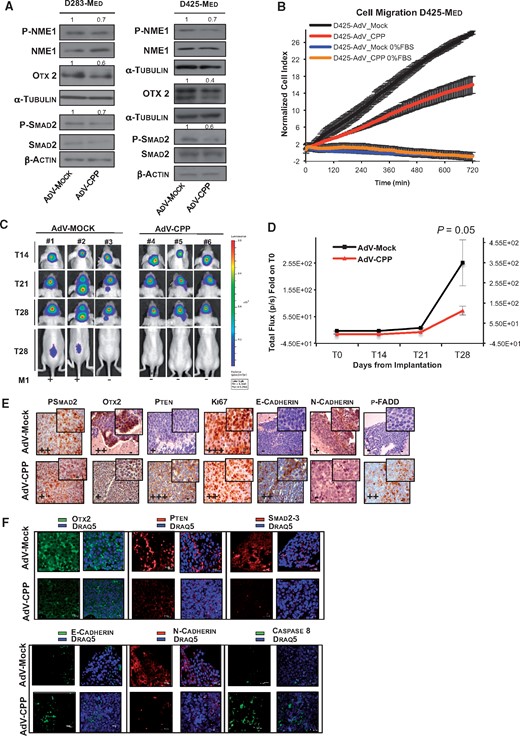
Competitive permeable peptide treatment impairs tumour progression and metastasis in vivo in xenograft models of MBGroup3. (A) Representative western blot with indicated antibodies of protein lysates from two MBGroup3 cell lines (D283-Med and D425-Med cells) infected with adenovirus particles with peptide sequence carrying the region of interaction of NME1 with PRUNE1 (AdV-CPP; see Supplementary material) and with empty vector (Adv-Mock) as control. CPP reduced phosphorylation levels of phospho-(Ser120-122-125)-NME1 (0.7-fold), phospho-SMAD2 (D283-Med, 0.7-fold; D425-Med, 0.6-fold) and OTX2 protein levels (D283-Med, 0.6-fold; D425-Med, 0.4-fold) in both MBGroup3 cell lines. (B) Normalized cell index as measure of cell migration of MBGroup3 D425-Med cells infected with AdV-CPP, or AdV-Mock as control, as generated by xCELLigence RTCA software. Migration kinetics were monitored in response to 10% FBS (red, black) and to 0% FBS (brown, blue) as negative controls. D425-Med cells infected with AdV-CPP (red) showed decreased migratory properties in response to 10% FBS gradient, in comparison with D425-Med cells infected with AdV-Mock (black). Data are means ± SD of triplicate samples. (C) Representative orthotopic xenograft experiment using MBGroup3 D425-Med cells stably expressing firefly Luciferase gene (D425-Luc cells) infected in vitro, with AdV-CPP or Adv-Mock as control, and then implanted into fourth right ventricle of cerebellum of six nude mice (three mice for each group). The mice were imaged every 7 days via in vivo bioluminescent imaging (IVIS Spectrum In Vivo Imaging System) to monitor tumour growth from time of implantation (T0) to 28 days after tumour implantation. (D) Tumour growth according to quantified photon emission (ph/s) from the region of interest of mice orthotopically injected with previously infected D425-Luc cells. The differences in total flux from the two groups of mice at 28 days from cell orthotopic implantation (T28) showed impairment of tumour growth in cerebellum in vivo in the mice implanted with AdV-CPP-infected D425-Med (P = 0.05; Supplementary Table 4). Data are means ± SD. (E and F) Representative immunohistochemistry (E) and indirect immunofluorescence staining (F) with indicated antibodies of paraffin-embedded cerebellar tumours generated by implanting D425-Luc cells into the fourth ventricle of nude mice (immunohistochemistry ×40; immunofluorescence ×63).
We also investigated CPP modulation of medulloblastoma growth in vivo with an orthotopic xenograft mouse model of MBGroup3, using D425-Med cells engineered to stably express the firefly Luciferase (Luc)gene (D425-Luc). Orthotopic implantation of these D425-Luc cells infected with AdV-CPP promoted significant decrease in primary tumour growth at 28 days post-implantation (P = 0.05; Fig. 4C, D and Supplementary Table 4). CPP also inhibited the metastatic dissemination in the spinal cord in those mice orthotopically injected with cells infected with AdV-CPP compared to controls (Fig. 4C). Both immunohistochemistry and immunofluorescence of primary tumour tissues from xenograft mice showed inhibition of the TGF-β pathway (i.e. decreased nuclear OTX2: 0.11-fold; nuclear phosphorylated SMAD2: 0.09-fold), upregulation of PTEN (10.11-fold), reduced EMT (i.e. N-cadherin loss: 0.14-fold; E-cadherin increase: 14.62-fold), and induction of apoptotic pathways [i.e. increased phosphorylated-[Ser194]-Fas-associated proteins with death domain (FADD) and Caspase-8 (8.47-fold); Fig. 4E, F and Supplementary Fig. 4E].
Altogether here, this demonstrated the contribution of the PRUNE1/NME1 protein complex to the activation of TGF-β signalling, which thus promotes cell migration and EMT through positive regulation of OTX2 in MBGroup3.
A small molecule inhibitor designed to target PRUNE1
Silencing endogenous PRUNE1 resulted in decreased proliferation of D283-Med and D425-Med cells (Supplementary Fig. 5A). Nuclear magnetic resonance (NMR) and molecular dynamic studies that defined the 3D structure of PRUNE1 allowed us to synthesize small molecules derived from the pyrimido-pyrimidine AA2.8 (Virgilio et al., 2012). A further modification then defined AA7.1 with increased solubility. NMR showed that PRUNE1 interacted structurally with AA7.1 through two amino acids, S365 and negatively charged D364, which significantly contributed to AA7.1 binding to C-term-PRUNE1 (Fig. 5A and B). Control studies using site-directed mutagenesis confirmed the binding affinity of AA7.1 for PRUNE1 (Supplementary Fig. 5B). Furthermore, the half maximal inhibitory concentration (IC50) of AA7.1 for the inhibition of cell proliferation index was also measured for both D283-Med and D425-Med cells, as 129.99 ± 52.05 µM (R = 0.97; data not shown).
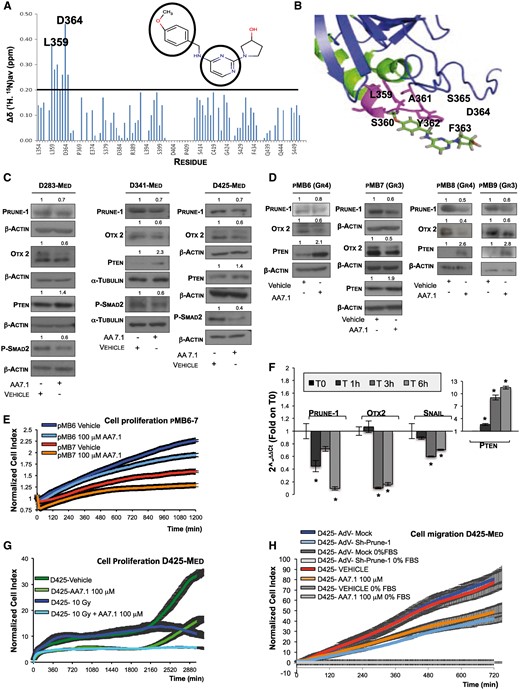
Effects of PRUNE1 silencing in MBGroup3 cells in vitro. (A) In vitro nuclear magnetic resonance protein–drug interaction studies. Variations in chemical shift in the (1H, 15N) heteronuclear single quantum coherence spectrum of 15N-labelled C-term-PRUNE1 domain upon formation of AA7.1–C-term-PRUNE1 complex. L365 and D364 amino acids were seen to significantly contribute to this complex formation. The chemical structure of AA7.1 is shown, together with the amino acid regions of interaction with saturation transfer difference signals (circles: protons of methoxyphenyl ring, protons of pyrimidine ring). (B) Amino acids that showed large variations upon complex formation with AA71 (magenta) mapped onto the C-term-PRUNE1 surface. (C) Representative western blots using protein lysates from indicated different MBGroup3 cell lines treated with 100 µM AA7.1 for 6 h or PBS as vehicle control. Densitometric values given showed downregulation of PRUNE1 (0.7-fold), similar in all AA7.1-treated medulloblastoma cells, compared to vehicle. AA7.1-treated MBGroup3 cells also showed decrease in phospho-Ser467-SMAD2 and OTX2 and upregulation of PTEN. β-Actin and α-tubulin levels were used as loading controls. (D) Representative western blots using indicated antibodies on cell lysates of adherent primary MBGroup3/4 cells (pMB7 and pMB6), and MBGroup3/4 medullospheres (pMB9 and pMB8) treated with 100 μM AA7.1 or PBS as vehicle for 6 h. Data showed reduction of PRUNE1 and OTX2 protein levels and increased PTEN protein levels in all AA7.1-treated primary MBGroup3 cells. β-Actin levels were used as the loading control. (E) Normalized cell index as measure of cell proliferation of primary MBGroup3 pMB7 and MBGroup4 pMB6 cells treated with AA7.1 (100 µM) or PBS as control, as generated by xCELLigence RTCA software. AA7.1 reduces the proliferation rate of both pMB6 (light-blue) and pMB7 (orange) compared to controls (blue and red, respectively). Data are means ± SD of samples assayed in triplicate. (F) Real-time PCR time-course expression analysis of PRUNE1, OTX2, SNAIL and PTEN mRNA levels in MBGroup3 D425-Med cells treated with 100 µM AA7.1 or PBS as control. Downregulation of PRUNE1 and increase in PTEN mRNA expression levels were seen in D425-Med cells at 1 h from addition of AA7.1, while OTX2 and SNAIL downregulation was seen after 3 h. Statistically significant reductions (*P < 0.05). (G) Normalized cell index as measure of cell proliferation of D425-Med cells treated with 100 µM AA7.1 or PBS as control, and irradiated with 10 Gy (at 1.2 Gy/min), as generated by xCELLigence RTCA software. D425-Med cells treated with AA7.1 and irradiated (light blue) showed decreased cell proliferation compared to those treated with AA7.1 (light green) or irradiated alone (blue). Data are means ± SD of samples assayed in triplicate. (H) Normalized cell index as measure of cell migration of MBGroup3 D425-Med cells infected with AdV-sh-PRUNE1 or AdV-Mock as control, or treated with AA7.1 (100 µM), as generated by xCELLigence RTCA software. Migration kinetics were monitored in response to 10% FBS. Both D425-Med cells infected with AdV-sh-PRUNE1 (light blue) or treated with AA7.1 (orange) showed decreased migratory properties in comparison with D425-Med cells infected with AdV-Mock (blue) or treated with PBS (red) as vehicle. Data are means ± SD of samples assayed in triplicate.
D283-Med, D425-Med and D341-Med cells were then treated with 100 µM AA7.1, which resulted in decreased endogenous PRUNE1 levels (Fig. 5C). This also affected the TGF-β cascade, with reduced SMAD2 phosphorylation and OTX2 expression, and increased PTEN expression (Fig. 5C). This was also shown in human primary adherent MBGroup3/4 cells (i.e. pMB6 and pMB7 cells) and in primary medullospheres derived from metastatic MBGroup3/4 patients (i.e. pMB9 and pMB8) (Fig. 5D). Furthermore, the treatment with AA7.1 of primary adherent MBGroup3/4 cells (i.e. pMB6 and pMB7 cells) resulted in inhibition of cell proliferation (Fig. 5E). Thus, overall, these data indicated that AA7.1 treatment of several cellular models of metastatic medulloblastoma affected the PRUNE1/PTEN axis and the cell proliferation.
To dissect out the mechanism of action of AA7.1, D425-Med cells were co-treated with AA7.1 and the protein synthesis inhibitor Cycloheximide, resulting in PRUNE1 accumulation. Thus, this AA7.1 treatment led to increased PRUNE1 degradation (Supplementary Fig. 5C). Then, time course expression analysis of D425-Med cells was carried out at defined intervals (1, 3 and 6 h), to monitor the gene targets previously identified (PRUNE1, OTX2, SNAIL, PTEN) when 100 µM AA7.1 was used to decrease PRUNE1 mRNA expression (0.44-fold at 1 h). Here there was early upregulation of PTEN (1 h: 2.63-fold), with inhibition of OTX2 (0.1-fold) and SNAIL (0.59-fold) at 3 h treatment. This inhibition of PRUNE1, OTX2 and SNAIL expression levels and increased PTEN mRNA levels were shown also after 6 h treatment (PRUNE1, 0.09-fold; OTX2, 0.16-fold; SNAIL, 0.7-fold; PTEN, 11.4-fold; Fig. 5F and Supplementary Table 1). Although these data only monitored RNA expression levels, they further indicated that it is possible to define these targets of AA7.1 inhibition. In addition, an alternative mechanism of PRUNE1-mediated PTEN upregulation independent of SNAIL expression can be postulated. Further studies will dissect out this particular hypothesis.
Combinatorial radiotherapy and chemotherapy is the standard protocol for the treatment of patients with metastatic medulloblastoma. To investigate this, we investigated whether AA7.1 in combination with radiotherapy might have future therapeutic implications. Cell proliferation rates were measured using electrical impedance of D425-Med cells pretreated with 100 µM AA7.1. These cells were also irradiated as a single dose of 10 Gy (at 1.2 Gy/min) prior to the proliferation experiment (Rieken et al., 2015). This combinatorial approach indeed decreased the proliferation rate of D425-Med cells (Fig. 5G).
Then, the pharmacological inhibition of PRUNE1 and its silencing were compared in D425-Med cells, followed by cell migration assay. Adenoviral silencing and pharmacological inhibition of PRUNE1 showed similar rates of inhibition of cell migration, measured as mean cell-index impedance (Fig. 5H). Similar results were obtained in D283-Med cells (data not shown). Overall, this thus confirms the data presented (Figs 3F and 5C). Most importantly, there was an increased fraction of apoptotic cells (75%) in the analysis of Annexin V and propidium iodide staining of D425-Med cells infected with AdV-sh-PRUNE1, compared to those treated with AA7.1 (Supplementary Fig. 5D). For the above reasons, the adenoviral-carrying sh-PRUNE1 strategy requires further in vivo analysis due to the substantial enhancement of apoptosis.
AA7.1 impairs medulloblastoma growth and metastases in vivo through targeting the PRUNE1/TGF-β/OTX2/PTEN axis
Toxicity studies were carried out with escalating-dose of AA7.1 in vivo in wild-type Balb/C mice (4 weeks old; grouped according to weight), as 15, 30 and 60 mg/kg intraperitoneally, administered daily for 1 week. These showed no immediate toxicity effects of AA7.1. Blood was collected and haematological and biochemical analyses were carried out, along with morphological evaluation of their organs (i.e. spleen, liver, kidneys). There were no significant variations in the haematological parameters (e.g. white blood cell count), and no signs of hepatotoxicity or nephrotoxicity across these three groups of mice, according to glutamate-pyruvate transaminase 1, glutamic oxaloacetic transaminase, creatinine blood levels, and blood urea nitrogen (Supplementary Table 5). The modulation of medulloblastoma growth and metastases by AA7.1 was then investigated in vivo using orthotopic xenograft mouse models of MBGroup3 and D425-Luc cells (Supplementary material). The maximum dose of AA7.1 was limited to 60 mg/kg intraperitoneally every 2 days (equivalent to 30 mg/kg/day) because of the borderline platelet counts (591.5 versus 610.0 × 109/l) in those mice treated with 60 mg/kg/daily (Supplementary Table 5), to prevent blood clots and thrombosis over long periods of treatment.
In this in vivo preclinical trial, after the tumours were established in the fourth ventricle (14 days post-injection), intraperitoneal administration of AA7.1 was started (60 mg/kg every 2 days; Fig. 6A). These data showed significantly decreased intracranial tumour growth after 1 week and 2 weeks of treatment for the AA7.1-treated mice (n = 6) compared to the vehicle group (n = 6) (T21, P = 0.01; T28, P < 0.00001; Fig. 6B). Here, AA7.1 impaired MBGroup3 cancer progression and also impaired metastasis to the spinal cord (Fig. 6A).
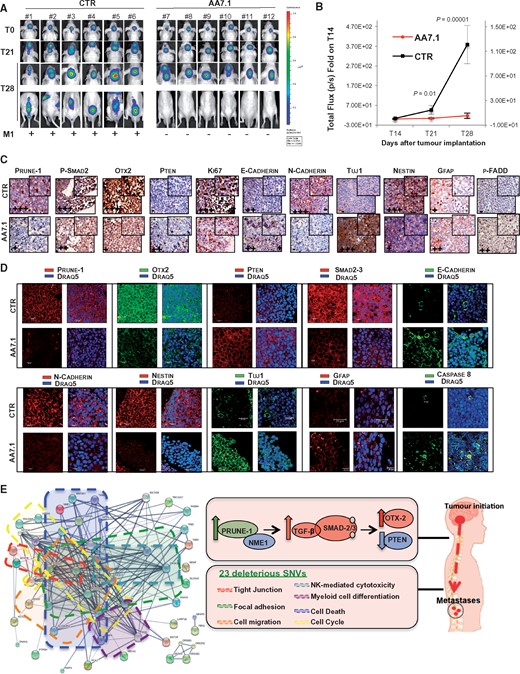
AA7.1 treatment impairs tumour progression and metastasis in vivo in models of MBGroup3. (A) Representative orthotopic xenograft experiment following injection into the fourth ventricle of nude mice (n = 12) of MBGroup3 D425-Luc cells. The mice were imaged every 7 days for in vivo bioluminescence acquisition, to monitor tumour growth. (B) Tumour growth according to quantified photon emission (ph/s) from the region of interest of mice orthotopically injected with D425-Luc cells treated in vivo with AA7.1 (60 mg/kg every 2 days, intraperitoneally) or with PBS as vehicle. Data are means ± SD. The differences in total flux from the two groups of mice at 21 days from cell orthotopic implantation (T21; after 7 days of AA7.1) and 28 days (T28; after 14 days of AA7.1) showed impairment of tumour growth in cerebellum in vivo in the AA7.1-treated mice (T21, P = 0.01; T28, P = 0.00001). (C and D) Representative immunohistochemistry (C) and indirect immunofluorescence staining (D) with indicated antibodies of paraffin-embedded cerebellar tumours generated by implanting D425-Luc cells into the fourth ventricle of nude mice (immunohistochemistry ×40; immunofluorescence ×63). (E) ‘Pro-metastatic’ protein network generated via Search Tool for Retrieval of Interacting Genes/Proteins (STRING) database using deleterious homozygous NSVs common to pMB6 and pMB7 (MBGroup4 and MBGroup3, respectively; see Supplementary material). Twenty-three deleterious mutations (Table 2), together with the PRUNE1-driven axis, take part in networks characterized by proteins involved in tight junctions (red dashed box), focal adhesions (green dashed box), regulation of cell migration (orange dashed box), regulation of cell death (blue dashed box), regulation of cell cycle (yellow dashed box), natural-killer-cell-mediated cytotoxicity (light blue dashed box), and positive regulation of myeloid leukocyte differentiation (purple dashed box). The central boxes and right diagram summarize the proposed mechanism by which PRUNE1 enhances canonical TGF-β signalling activation through the binding to NME1, which thus enhances OTX2 upregulation and PTEN inhibition in metastatic MBGroup3, as dissected out from the different cell models of MBGroup3. The pathways affected by the 23 deleterious homozygous SNVs common to primary human metastatic pMB6 and pMB7 (MBGroup4, MBGroup3, respectively) are also indicated. Together with the newly identified PRUNE1-driven axis, these variants take part in this pro-metastatic interactions network.
Immunohistochemistry and immunofluorescence of the cerebellar tumours in these mice treated with AA7.1 in vivo showed reduced PRUNE1 levels (0.44-fold), inhibition of nuclear phosphorylated SMAD2 (0.6-fold) and OTX2 (0.2-fold), and overexpression of PTEN (3.8-fold). In the same tissues, the treated mice also showed inhibition of EMT with N-cadherin loss (0.29-fold) and increased E-cadherin (2.98-fold). Moreover, there was reduction of cell proliferative index (as decrease of Ki-67-positive tumour cells), together with inhibition of neural stem/progenitor cell Nestin (0.18-fold) and increased neuronal differentiation marker neuron-specific class III β-tubulin Tuj1 (2.84-fold). Nestin is known to define novel progenitor cells of cerebellar granule neurons, and thus to generate propagating tumorigenic medulloblastoma (Li et al., 2013). Altogether, these demonstrated neuronal differentiation processes in these AA7.1-treated mice. Here, there was also upregulation of GFAP in tumours from these AA7.1-treated mice (2.85-fold). This thus suggested that the pharmacological inhibition of PRUNE1 can increase the levels of GFAP-positive cells, which are known to have a positive prognostic significance in medulloblastoma (Goldberg-Stern et al., 1991). These data were not observed in the trial with CPP (data not shown), showing an alternative unknown mechanism of regulation. In addition, there was induction of pro-apoptotic processes in the tumour tissues derived from the AA7.1-treated mice, which included increased phosphorylated-FADD and Caspase-8 (2.08-fold; Fig. 6C, D and Supplementary Fig. 5E).
Of note, AA7.1 did not show any signs of toxicity in these treated mice, as demonstrated by no decrease in total body weight in the treated mice compared to controls (data not shown) and by the haematological parameters (Supplementary Table 5). There was also an absence of drug-induced hepatotoxicity and renal failure, as seen by glutamate-pyruvate transaminase 1, glutamic oxaloacetic transaminase and creatinine blood levels, and blood urea nitrogen from three tumour-bearing mice (Supplementary Table 5). These data are of importance, because at primary sites there was inhibition of cell proliferation by AA7.1 treatment.
Discussion
The molecular mechanisms that drive medulloblastoma metastasis and the disease features within the relapse setting are currently not known. The present study has revealed a role for PRUNE1 in metastatic MBGroup3.
We have shown that PRUNE1 positively enhances canonical TGF-β signalling leading to enhancement of SNAIL expression and downregulation of PTEN. It was previously reported that subtle reductions in PTEN can have a profound impact on tumour susceptibility and progression (Alimonti et al., 2010). Immunohistopathological data (Fig. 3G and Supplementary Fig. 4B) indicated that ‘PRUNE1high/PTENlow’ is independent of c-MYC amplification, which thus supports the concept of two phenotypes in MBGroup3 (i.e. PRUNE1high/PTENlow, and vice versa), with different correlations to prognosis and metastatic events. As such, these two subcategories of MBGroup3 (i.e. PTENhigh, PTENlow) should follow different treatment regimens (von Bueren et al., 2016).
Among the genes with a role in neurogenesis or neural differentiation, GLI2 and CYFIP1 were also inversely correlated to PRUNE1 in medulloblastoma (Supplementary Table 2). We believe that regulation of GLI2 and CYFIP1 expression in MBGroup3-PRUNE1high tumours will strengthen the neural programme of differentiation, development and regulation of neurogenesis, which will be of therapeutic impact (see Fig. 6D: Tuj1/Nestin and GFAP target expression in the AA7.1-treated mice versus those not treated).
The in vitro data presented here using the AA7.1 inhibitor of PRUNE1 and irradiation (Fig. 5G) have promising indications for the treatment of MBGroup3 patients. As described, AA7.1 did not show toxicity in terms of haematological and biochemical functions in AA7.1-treated mice. An increase in white blood cell counts and lymphocytes was also seen in these AA7.1-treated mice (Supplementary Table 5). This suggests potential pharmacological enhancement of natural killer cells and B cells in these athymic Nude-Foxn1nu mice (i.e. deficient in T cells, but with normal B cells and natural killer cells). This phenomenon resembles TGF-β inhibition with enhancement of growth and maturation of multiple immune cell lineages, including T cells (Yang et al., 2010; Quatromoni et al., 2013). The action of AA7.1 within immune cell physiology needs future studies. At this time, we believe that other tumours of the posterior fossa, where regulation of TGF-β signalling might have a critical role, can also be treated with AA7.1.
As previously discussed, TGF-β signalling has a role in cell physiology in the immune system. These genes identified through NGS studies that retain deleterious loss-of-function mutations might also take part in the cross-talk between tumorigenic and immune cells in the tumour microenvironment. The data presented here include 23 deleterious NSVs in common with metastatic MBGroup3/4 primary tumour patients (Table 2). Of these, we applied further analysis using the Kyoto Encyclopaedia of Genes and Genomes (KEGG), to define a network of protein interactions (Fig. 6E). We showed that focal adhesion (pathway ID:5200), tight junction regulation (pathway ID:4530) and cell-mediated cytotoxicity of natural killer cells (pathway ID:4650) are new functions that are involved in metastatic spread, together with de-regulation of the innate immune response that involves natural killer cells (Lee et al., 2015). In the same protein network, there were proteins involved in regulation of cell migration (GO.0030334), cell-matrix adhesion (GO.0045785, GO.0022407, GO.0001952), regulation of neuron death (GO.1901214), cell-cycle control (GO.0051726), and regulation of myeloid leukocyte differentiation (GO.0002763, GO.0002761). Altogether, these NSVs in single genes generated a network with functions related to cell dissemination and immune control within metastatic medulloblastoma progression (Fig. 6E). For the above reasons, we envision the use with caution of check-point inhibitors in future therapies of metastatic MBGroup3/4.
Deleterious pro-metastatic ‘non-synonymous homozygous’ gene variants common to MBGroup3/4
| Gene symbol . | Location . | Allele . | Strand . | Uploaded variation . | cDNA position . | Protein position . | Amino acids . | Reference . |
|---|---|---|---|---|---|---|---|---|
| ABCA2 | 9:137018787–137018787 | T | −1 | rs908828 | 1801 | 584 | P/H | Mack et al., 2008 |
| ACTL9 | 19:8698550–8698550 | A | −1 | rs10410943 | 273 | 51 | A/V | Paternoster et al., 2011 |
| CHMP4A | 14:24210671–24210671 | T | −1 | rs2295322 | 887 | 196 | G/R | Tsang et al., 2006 |
| DNAH1 | 3:52396682–52396682 | A | 1 | rs365048 | 11756 | 3832 | R/H | Neesen et al., 2001 |
| FBN1 | 15:48515440–48515440 | C | −1 | rs4775765 | 1871 | 472 | Y/C | Sakai et al., 2016 |
| GNA15 | 19:3150240–3150240 | A | 1 | rs310680 | 698 | 147 | C/Y | Contos et al., 2002 |
| LARP1B | 4:128122049–128122049 | G | 1 | rs12508837 | 1596 | 462 | P/R | Stavraka and Blagden, 2015 |
| MST1R | 3:49891258–49891258 | T | −1 | rs7433231 | 3611 | 1195 | G/S | Dai et al., 2016b |
| MYH15 | 3:108470146–108470146 | A | −1 | rs9868484 | 1567 | 504 | H/Y | Stedman et al., 2004 |
| MYH7B | 20:34995528–34995528 | A | 1 | rs2425015 | 3111 | 1007 | E/K | van Rooij et al., 2009 |
| NEMF | 14:49832244–49832244 | A | −1 | rs3100906 | 1219 | 257 | S/C | Shao et al., 2015 |
| OPTN | 10:13124076–13124076 | A | 1 | rs523747 | 1326 | 322 | E/K | Slowicka et al., 2016 |
| OR2W3 | 1:247896174–247896174 | C | 1 | rs12139390 | 588 | 196 | E/D | Ma et al., 2015 |
| OR51B2 | 11:5323898–5323898 | A | −1 | rs10837814 | 400 | 134 | L/F | Solovieff et al., 2010 |
| OR52N2 | 11:5821126–5821126 | G | 1 | rs8181512 | 813 | 264 | H/R | Malnic et al., 2004 |
| OR56B1 | 11:5736832–5736832 | C | 1 | rs7397032 | 316 | 106 | C/R | Malnic et al., 2004 |
| PARP4 | 13:24455080–24455080 | T | −1 | rs2275660 | 2801 | 899 | A/T | Long et al., 2016 |
| PCK2 | 14:24098289–24098289 | A | 1 | rs3021119 | 630 | 121 | P/Q | Luo et al., 2017 |
| PTPRH | 19:55188086–55188086 | T | −1 | rs890870 | 2490 | 823 | E/K | Nagano et al., 2003 |
| RHOD | 11:67070494–67070494 | T | 1 | rs4930409 | 485 | 134 | R/C | Long and Simpson, 2017 |
| SLC5A3 | 21:34095346–34095346 | A | 1 | rs8129891 | 660 | 50 | A/T | Dai et al., 2016a |
| TAB3 | X:30854485–30854485 | A | −1 | rs5927629 | 1727 | 394 | R/W | Jin et al., 2004 |
| TBC1D17 | 19:49880334–49880334 | A | 1 | rs8109661 | 550 | 84 | G/D | Vaibhava et al., 2012 |
| Gene symbol . | Location . | Allele . | Strand . | Uploaded variation . | cDNA position . | Protein position . | Amino acids . | Reference . |
|---|---|---|---|---|---|---|---|---|
| ABCA2 | 9:137018787–137018787 | T | −1 | rs908828 | 1801 | 584 | P/H | Mack et al., 2008 |
| ACTL9 | 19:8698550–8698550 | A | −1 | rs10410943 | 273 | 51 | A/V | Paternoster et al., 2011 |
| CHMP4A | 14:24210671–24210671 | T | −1 | rs2295322 | 887 | 196 | G/R | Tsang et al., 2006 |
| DNAH1 | 3:52396682–52396682 | A | 1 | rs365048 | 11756 | 3832 | R/H | Neesen et al., 2001 |
| FBN1 | 15:48515440–48515440 | C | −1 | rs4775765 | 1871 | 472 | Y/C | Sakai et al., 2016 |
| GNA15 | 19:3150240–3150240 | A | 1 | rs310680 | 698 | 147 | C/Y | Contos et al., 2002 |
| LARP1B | 4:128122049–128122049 | G | 1 | rs12508837 | 1596 | 462 | P/R | Stavraka and Blagden, 2015 |
| MST1R | 3:49891258–49891258 | T | −1 | rs7433231 | 3611 | 1195 | G/S | Dai et al., 2016b |
| MYH15 | 3:108470146–108470146 | A | −1 | rs9868484 | 1567 | 504 | H/Y | Stedman et al., 2004 |
| MYH7B | 20:34995528–34995528 | A | 1 | rs2425015 | 3111 | 1007 | E/K | van Rooij et al., 2009 |
| NEMF | 14:49832244–49832244 | A | −1 | rs3100906 | 1219 | 257 | S/C | Shao et al., 2015 |
| OPTN | 10:13124076–13124076 | A | 1 | rs523747 | 1326 | 322 | E/K | Slowicka et al., 2016 |
| OR2W3 | 1:247896174–247896174 | C | 1 | rs12139390 | 588 | 196 | E/D | Ma et al., 2015 |
| OR51B2 | 11:5323898–5323898 | A | −1 | rs10837814 | 400 | 134 | L/F | Solovieff et al., 2010 |
| OR52N2 | 11:5821126–5821126 | G | 1 | rs8181512 | 813 | 264 | H/R | Malnic et al., 2004 |
| OR56B1 | 11:5736832–5736832 | C | 1 | rs7397032 | 316 | 106 | C/R | Malnic et al., 2004 |
| PARP4 | 13:24455080–24455080 | T | −1 | rs2275660 | 2801 | 899 | A/T | Long et al., 2016 |
| PCK2 | 14:24098289–24098289 | A | 1 | rs3021119 | 630 | 121 | P/Q | Luo et al., 2017 |
| PTPRH | 19:55188086–55188086 | T | −1 | rs890870 | 2490 | 823 | E/K | Nagano et al., 2003 |
| RHOD | 11:67070494–67070494 | T | 1 | rs4930409 | 485 | 134 | R/C | Long and Simpson, 2017 |
| SLC5A3 | 21:34095346–34095346 | A | 1 | rs8129891 | 660 | 50 | A/T | Dai et al., 2016a |
| TAB3 | X:30854485–30854485 | A | −1 | rs5927629 | 1727 | 394 | R/W | Jin et al., 2004 |
| TBC1D17 | 19:49880334–49880334 | A | 1 | rs8109661 | 550 | 84 | G/D | Vaibhava et al., 2012 |
Details of the 23 deleterious ‘non-synonymous homozygous’ gene variants common to metastatic MBGroup3-4 primary cells (pMB6, pMB7 cells), as identified by next generation sequencing using whole exome sequencing technology. The selected variants involved in metastatic processes in MBGroup3-4 identify a ‘pro-metastatic’ protein network (related to Fig. 6E).
Deleterious pro-metastatic ‘non-synonymous homozygous’ gene variants common to MBGroup3/4
| Gene symbol . | Location . | Allele . | Strand . | Uploaded variation . | cDNA position . | Protein position . | Amino acids . | Reference . |
|---|---|---|---|---|---|---|---|---|
| ABCA2 | 9:137018787–137018787 | T | −1 | rs908828 | 1801 | 584 | P/H | Mack et al., 2008 |
| ACTL9 | 19:8698550–8698550 | A | −1 | rs10410943 | 273 | 51 | A/V | Paternoster et al., 2011 |
| CHMP4A | 14:24210671–24210671 | T | −1 | rs2295322 | 887 | 196 | G/R | Tsang et al., 2006 |
| DNAH1 | 3:52396682–52396682 | A | 1 | rs365048 | 11756 | 3832 | R/H | Neesen et al., 2001 |
| FBN1 | 15:48515440–48515440 | C | −1 | rs4775765 | 1871 | 472 | Y/C | Sakai et al., 2016 |
| GNA15 | 19:3150240–3150240 | A | 1 | rs310680 | 698 | 147 | C/Y | Contos et al., 2002 |
| LARP1B | 4:128122049–128122049 | G | 1 | rs12508837 | 1596 | 462 | P/R | Stavraka and Blagden, 2015 |
| MST1R | 3:49891258–49891258 | T | −1 | rs7433231 | 3611 | 1195 | G/S | Dai et al., 2016b |
| MYH15 | 3:108470146–108470146 | A | −1 | rs9868484 | 1567 | 504 | H/Y | Stedman et al., 2004 |
| MYH7B | 20:34995528–34995528 | A | 1 | rs2425015 | 3111 | 1007 | E/K | van Rooij et al., 2009 |
| NEMF | 14:49832244–49832244 | A | −1 | rs3100906 | 1219 | 257 | S/C | Shao et al., 2015 |
| OPTN | 10:13124076–13124076 | A | 1 | rs523747 | 1326 | 322 | E/K | Slowicka et al., 2016 |
| OR2W3 | 1:247896174–247896174 | C | 1 | rs12139390 | 588 | 196 | E/D | Ma et al., 2015 |
| OR51B2 | 11:5323898–5323898 | A | −1 | rs10837814 | 400 | 134 | L/F | Solovieff et al., 2010 |
| OR52N2 | 11:5821126–5821126 | G | 1 | rs8181512 | 813 | 264 | H/R | Malnic et al., 2004 |
| OR56B1 | 11:5736832–5736832 | C | 1 | rs7397032 | 316 | 106 | C/R | Malnic et al., 2004 |
| PARP4 | 13:24455080–24455080 | T | −1 | rs2275660 | 2801 | 899 | A/T | Long et al., 2016 |
| PCK2 | 14:24098289–24098289 | A | 1 | rs3021119 | 630 | 121 | P/Q | Luo et al., 2017 |
| PTPRH | 19:55188086–55188086 | T | −1 | rs890870 | 2490 | 823 | E/K | Nagano et al., 2003 |
| RHOD | 11:67070494–67070494 | T | 1 | rs4930409 | 485 | 134 | R/C | Long and Simpson, 2017 |
| SLC5A3 | 21:34095346–34095346 | A | 1 | rs8129891 | 660 | 50 | A/T | Dai et al., 2016a |
| TAB3 | X:30854485–30854485 | A | −1 | rs5927629 | 1727 | 394 | R/W | Jin et al., 2004 |
| TBC1D17 | 19:49880334–49880334 | A | 1 | rs8109661 | 550 | 84 | G/D | Vaibhava et al., 2012 |
| Gene symbol . | Location . | Allele . | Strand . | Uploaded variation . | cDNA position . | Protein position . | Amino acids . | Reference . |
|---|---|---|---|---|---|---|---|---|
| ABCA2 | 9:137018787–137018787 | T | −1 | rs908828 | 1801 | 584 | P/H | Mack et al., 2008 |
| ACTL9 | 19:8698550–8698550 | A | −1 | rs10410943 | 273 | 51 | A/V | Paternoster et al., 2011 |
| CHMP4A | 14:24210671–24210671 | T | −1 | rs2295322 | 887 | 196 | G/R | Tsang et al., 2006 |
| DNAH1 | 3:52396682–52396682 | A | 1 | rs365048 | 11756 | 3832 | R/H | Neesen et al., 2001 |
| FBN1 | 15:48515440–48515440 | C | −1 | rs4775765 | 1871 | 472 | Y/C | Sakai et al., 2016 |
| GNA15 | 19:3150240–3150240 | A | 1 | rs310680 | 698 | 147 | C/Y | Contos et al., 2002 |
| LARP1B | 4:128122049–128122049 | G | 1 | rs12508837 | 1596 | 462 | P/R | Stavraka and Blagden, 2015 |
| MST1R | 3:49891258–49891258 | T | −1 | rs7433231 | 3611 | 1195 | G/S | Dai et al., 2016b |
| MYH15 | 3:108470146–108470146 | A | −1 | rs9868484 | 1567 | 504 | H/Y | Stedman et al., 2004 |
| MYH7B | 20:34995528–34995528 | A | 1 | rs2425015 | 3111 | 1007 | E/K | van Rooij et al., 2009 |
| NEMF | 14:49832244–49832244 | A | −1 | rs3100906 | 1219 | 257 | S/C | Shao et al., 2015 |
| OPTN | 10:13124076–13124076 | A | 1 | rs523747 | 1326 | 322 | E/K | Slowicka et al., 2016 |
| OR2W3 | 1:247896174–247896174 | C | 1 | rs12139390 | 588 | 196 | E/D | Ma et al., 2015 |
| OR51B2 | 11:5323898–5323898 | A | −1 | rs10837814 | 400 | 134 | L/F | Solovieff et al., 2010 |
| OR52N2 | 11:5821126–5821126 | G | 1 | rs8181512 | 813 | 264 | H/R | Malnic et al., 2004 |
| OR56B1 | 11:5736832–5736832 | C | 1 | rs7397032 | 316 | 106 | C/R | Malnic et al., 2004 |
| PARP4 | 13:24455080–24455080 | T | −1 | rs2275660 | 2801 | 899 | A/T | Long et al., 2016 |
| PCK2 | 14:24098289–24098289 | A | 1 | rs3021119 | 630 | 121 | P/Q | Luo et al., 2017 |
| PTPRH | 19:55188086–55188086 | T | −1 | rs890870 | 2490 | 823 | E/K | Nagano et al., 2003 |
| RHOD | 11:67070494–67070494 | T | 1 | rs4930409 | 485 | 134 | R/C | Long and Simpson, 2017 |
| SLC5A3 | 21:34095346–34095346 | A | 1 | rs8129891 | 660 | 50 | A/T | Dai et al., 2016a |
| TAB3 | X:30854485–30854485 | A | −1 | rs5927629 | 1727 | 394 | R/W | Jin et al., 2004 |
| TBC1D17 | 19:49880334–49880334 | A | 1 | rs8109661 | 550 | 84 | G/D | Vaibhava et al., 2012 |
Details of the 23 deleterious ‘non-synonymous homozygous’ gene variants common to metastatic MBGroup3-4 primary cells (pMB6, pMB7 cells), as identified by next generation sequencing using whole exome sequencing technology. The selected variants involved in metastatic processes in MBGroup3-4 identify a ‘pro-metastatic’ protein network (related to Fig. 6E).
In summary, the present study highlights the function of PRUNE1/NME1 and dissects out a novel PRUNE1/NME1/TGF-β/OTX2/PTEN axis in metastatic MBGroup3. This ‘metastatic’ axis, together with the novel identified ‘loss-of-function’ mutations, is responsible for initiation of a pro-migratory phenotype of MBgroup3 tumours, which can then result in metastastatic dissemination (Fig. 6E). These data have allowed the identification of several new metastasis-suppressing gene drivers that represent excellent new potential targets for therapeutic treatments of metastatic medulloblastoma.
Acknowledgements
The authors thank Dr. J.Wrana, Dr. J.Massague Addgene and Dr. B.Volgestein as providers of plasmids (Addgene, #11742, #14930, #16495). The authors dedicate this study to the parents of Leonardo Andrini, a child affected by MBGroup3 who died in October 2015.
Funding
We thank the following Resources Agencies for grant support: Fondazione Adolfo Volpe e Associazione Pediatri di Famiglia, EU-FP7-TUMIC-HEALTH-F2-2008-2016662 (MZ), the Italian Association for Cancer research (AIRC) Grant IG # 11963 (MZ), the Regione Campania L.g.R: N.5 (MZ), the European National Funds PON01-02388/1 2007-2013 (MZ), POR Rete delle Biotecnologie in Campania Movie (MZ), the European School of Molecular Medicine (SEMM) for the felloship (VF), the iCARE International AIRC felllowship (PDA) and the DMMBM Federico II fellowship (FPP).
Supplementary material
Supplementary material is available at Brain online.
Abbreviations
- CPP
competitive permeable peptide
- EMT
epithelial–mesenchymal transition
- MBGroup3/4
medulloblastoma group 3/4
References
Author notes
Veronica Ferrucci and Pasqualino de Antonellis authors contributed equally to this work.


![PRUNE1 expression in medulloblastoma. (A and B) RNA log2 expression of PRUNE1 derived from transcriptome analysis from the primary cohort of medulloblastoma (A; Boston cohort; n = 187; P < 0.05) and from the publicly available dataset of medulloblastoma (B; Cavalli; n = 763; P < 0.001), grouped according to the molecular and subtype disease variants. PRUNE1 was highly expressed in MBGroup3 and MBGroup4. Normal CB = normal cerebellum. (C) RNA log2 expression of PRUNE1 across medulloblastoma primary tumours [i.e. M1 (n = 176) versus M0 (n = 397)] in the publicly available Cavalli dataset (n = 573 tumours). There were higher PRUNE1 expression levels in metastatic M1 compared to M0 samples (P = 0.05). (D) Real-time PCR analysis on a ‘non-overlapping’ cohort of our medulloblastoma tissues from Italy, according to metastatic status at diagnosis (M0, n = 26; M1, n = 8). PRUNE1 expression levels were higher in metastatic M1 tumours compared to M0 (P = 0.05). See Supplementary Table 1 for real-time PCR expression data. (E) Kaplan-Meyer analysis for event-free survival (EFS) of MBGroup3 and MBGroup4 patients according to PRUNE1 expression levels. Patients who showed higher levels of PRUNE1 expression (n = 6) showed significantly shorter event-free survival compared to those with lower levels of PRUNE1 expression (n = 4) (P < 0.05). See Supplementary Table 1 for real-time PCR expression data. (F) RNA log2 expression of NME1 across the primary medulloblastoma samples from the publically available Cavalli dataset (n = 763; P < 0.001) grouped according to molecular subtypes. NME1 was highly expressed in MBGroup3-γ subtype.](https://oup.silverchair-cdn.com/oup/backfile/Content_public/Journal/brain/141/5/10.1093_brain_awy039/1/m_awy039f1.jpeg?Expires=1716357325&Signature=cXPNQU98uU3JLWa6s81nj-zwvdFtHgRcZZ3PJ6O8-BCEWtD72LN3F24jZP45dxIvlFJ9~aR0Fi-XvKTf1~2fABiRecpCknJBvU-1XIjcgi9jUjf-mZk6oxlnvBL3eNH8MI2KMuRNrqqeuiQTMw0seEvRSvyht5mX4l88oY0FgzYLpIWalvOTZCmNABgcmiTSkqkWcVAinH9Y7eUKHkZfpAB3ai5srvkL7vFYeaijd-ohkzAReCexs4j~k~ZC3yaSqhyrh4gyqY-IV8Kn4KOdGtDjEXweLQ00JopVEzyiSUPrpEa-6UXZ133bZ~DKUFulZBsOiBPjlTgMy~scfce1WA__&Key-Pair-Id=APKAIE5G5CRDK6RD3PGA)