Abstract
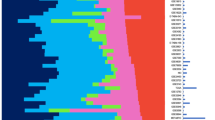
A better molecular characterization of breast cell lines (BCL) may help discover new markers to apply to tumour samples. We performed gene and protein expression profiling of 31 BCL using whole-genome DNA microarrays and immunohistochemistry (IHC) on ‘cell microarrays’ (CMA), respectively. Global hierarchical clustering discriminated two groups of BCL: group I corresponded to luminal cell lines, group II to basal and mesenchymal cell lines. Correlations with centroids calculated from a published ‘intrinsic 500-gene set’ assigned 15 cell lines as luminal, eight as basal and four as mesenchymal. A set of 1.233 genes was differentially expressed between basal and luminal samples. Mesenchymal and basal subtypes were rather similar and discriminated by only 227 genes. The expression of 10 proteins (CAV1, CD44, EGFR, MET, ETS1, GATA3, luminal cytokeratin CK19, basal cytokeratin CK5/6, CD10, and ERM protein moesin) encoded by luminal vs basal discriminator genes confirmed the subtype classification and the validity of the identified markers. Our BCL basal/luminal signature correctly re-classified the published series of tumour samples that originally served to identify the molecular subtypes, suggesting that the identified markers should be useful for tumour classification and might represent promising targets for disease management.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Abd El-Rehim DM, Ball G, Pinder SE, Rakha E, Paish C, Robertson JF et al. (2005). Int J Cancer 116: 340–350.
Al-Hajj M, Wicha MS, Benito-Hernandez A, Morrison SJ, Clarke MF . (2003). PNAS 100: 3983–3988.
Bertotti A, Comoglio PM . (2003). Trends Biochem Sci 28: 527–533.
Bertucci F, Borie N, Ginestier C, Groulet A, Charafe-Jauffret E, Adelaide J et al. (2004a). Oncogene 23: 2564–2575.
Bertucci F, Finetti P, Rougemont J, Charafe-Jauffret E, Cervera N, Tarpin C et al. (2005). Cancer Res 65: 2170–2178.
Bertucci F, Finetti P, Rougemont J, Charafe-Jauffret E, Nasser V, Loriod B et al. (2004b). Cancer Res 64: 8558–8565.
Bertucci F, Houlgatte R, Benziane A, Granjeaud S, Adelaide J, Tagett R et al. (2000). Hum Mol Genet 9: 2981–2991.
Birnbaum D, Bertucci F, Ginestier C, Tagett R, Jacquemier J, Charafe-Jauffret E . (2004). Int J Oncol 25: 249–258.
Boecker W, Buerger H . (2003). Cell Prolif 36 (Suppl 1): 73–84.
Carmeci C, Thompson DA, Kuang WW, Lightdale N, Furthmayr H, Weigel RJ . (1998). Surgery 124: 211–217.
Chang H, Sneddon J, Alizadeh A, Sood R, West R, Montgomery K et al. (2004). PLOS Biology 2: 206–214.
Chang JC, Wooten EC, Tsimelzon A, Hilsenbeck SG, Gutierrez MC, Elledge R et al. (2003). Lancet 362: 362–369.
Christensen JG, Burrows J, Salgia R . (2005). Cancer Lett 225: 1–26.
Clarke CL, Sandle J, Parry SC, Reis-Filho JS, O'Hare MJ, Lakhani SR . (2004). J Pathol 204: 147–152.
Condeelis J, Singer R, Segall JE . (2005). Ann Rev Cell Dev Biol 21: 695–718.
Dontu G, Al-Hajj M, Abdallah WM, Clarke MF, Wicha MS . (2003). Cell Prolif 36 (Suppl 1): 59–72.
Ediger TR, Park SE, Katzenellenbogen BS . (2002). Mol Endocrinol 16: 1828–1839.
Eisen MB, Spellman PT, Brown PO, Botstein D . (1998). Proc Natl Acad Sci USA 95: 14863–14868.
Ethier SP, Mahacek ML, Gullick WJ, Frank TS, Weber BL . (1993). Cancer Res 53: 627–635.
Farmer P, Bonnefoi H, Becette V, Tubiana-Hulin M, Fumoleau P, Larsimont D et al. (2005). Oncogene 24: 4660–4671.
Forozan F, Veldman R, Ammerman CA, Parsa NZ, Kallioniemi A, Kallioniemi OP et al. (1999). Br J Cancer 81: 1328–1334.
Foulkes WD, Brunet JS, Stefansson IM, Straume O, Chappuis PO, Begin LR et al. (2004). Cancer Res 64: 830–835.
Foulkes WD, Stefansson IM, Chappuis PO, Begin LR, Goffin JR, Wong N et al. (2003). J Natl Cancer Inst 95: 1482–1485.
Ginestier C, Charafe-Jauffret E, Bertucci F, Eisinger F, Geneix J, Bechlian D et al. (2002). Am J Pathol 161: 1223–1233.
Golub TR, Slonim DK, Tamayo P, Huard C, Gaasenbeek M, Mesirov JP et al. (1999). Science 286: 531–537.
Hackett AJ, Smith HS, Springer EL, Owens RB, Nelson-Rees WA, Riggs JL et al. (1977). J Natl Cancer Inst 58: 1795–1806.
Hedenfalk I, Ringner M, Ben-Dor A, Yakhini Z, Chen Y, Chebil G et al. (2003). Proc Natl Acad Sci USA 100: 2532–2537.
Irizarry RA, Hobbs B, Collin F, Beazer-Barclay YD, Antonellis KJ, Scherf U et al. (2003). Biostatistics 4: 249–264.
Jacquemier J, Ginestier C, Rougemont J, Bardou V-J, Charafe-Jauffret E, Geneix J et al. (2005). Cancer Research 65: 767–779.
Lacroix M, Leclercq G . (2004). Breast Cancer Res Treat 83: 249–289.
Leibl S, Gogg-Kammerer M, Sommersacher A, Denk H, Moinfar F . (2005). Am J Surg Pathol 29: 347–353.
Lengyel E, Prechtel D, Resau JH, Gauger K, Welk A, Lindemann K et al. (2005). Int J Cancer 113: 678–682.
Martin TA, Harrison G, Mansel RE, Jiang WG . (2003). Crit Rev Oncol Hematol 46: 165–186.
Mobus VJ, Moll R, Gerharz CD, Kieback DG, Merk O, Runnebaum IB et al. (1998a). Int J Cancer 77: 415–423.
Mobus VJ, Moll R, Gerharz CD, Kieback DG, Merk O, Runnebaum IB et al. (1998b). Int J Cancer 77: 415–423.
Nakamura N, Oshiro N, Fukata Y, Amano M, Fukata M, Kuroda S et al. (2000). Genes Cells 5: 571–581.
Navarro A, nand-Apte B, Parat MO . (2004). FASEB J 18: 1801–1811.
Nugoli M, Chuchana P, Vendrell J, Orsetti B, Ursule L, Nguyen C et al. (2003). BMC Cancer 3: 13–17.
Parr C, Watkins G, Mansel RE, Jiang WG . (2004). Clin Cancer Res 10: 202–211.
Paumelle R, Tulasne D, Kherrouche Z, Plaza S, Leroy C, Reveneau S et al. (2002). Oncogene 21: 2309–2319.
Perou CM, Sorlie T, Eisen MB, van de RM, Jeffrey SS, Rees CA et al. (2000). Nature 406: 747–752.
Razandi M, Oh P, Pedram A, Schnitzer J, Levin ER . (2002). Mol Endocrinol 16: 100–115.
Ross DT, Perou CM . (2001). Dis Markers 17: 99–109.
Sangiorgio V, Pitto M, Palestini P, Masserini M . (2004). Ital J Biochem 53: 98–111.
Schindelmann S, Windisch J, Grundmann R, Kreienberg R, Zeillinger R, Deissler H . (2002). Tumor Biol 23: 139–145.
Sorlie T, Perou CM, Tibshirani R, Aas T, Geisler S, Johnsen H et al. (2001). Proc Natl Acad Sci USA 98: 10869–10874.
Sorlie T, Tibshirani R, Parker J, Hastie T, Marron JS, Nobel A et al. (2003). Proc Natl Acad Sci USA 100: 8418–8423.
Sprenger RR, Speijer D, Back JW, De Koster CG, Pannekoek H, Horrevoets AJ . (2004). Electrophoresis 25: 156–172.
Stingl J, Eaves CJ, Zandieh I, Emerman JT . (2001). Breast Cancer Res Treat 67: 93–109.
Theillet C, Adélaide J, Louason G, Bonnet-Dorion F, Jacquemier J, Adnane J et al. (1993). Genes Chromosomes Cancer 7: 219–226.
Tomlinson GE, Chen TT, Stastny VA, Virmani AK, Spillman MA, Tonk V et al. (1998). Cancer Res 58: 3237–3242.
Troester MA, Hoadley KA, Sorlie T, Herbert BS, Borresen-Dale AL, Lonning PE et al. (2004). Cancer Res 64: 4218–4226.
Trusolino L, Comoglio PM . (2002). Nat Rev Cancer 2: 289–300.
Walker JA, Carder PJ . (2003). Histopathology 42: 300–301.
Welm AL, Kim S, Welm BE, Bishop JM . (2005). Proc Natl Acad Sci USA 102: 4324–4329.
Williams TM, Lisanti MP . (2005). Am J Physiol Cell Physiol 288: C494–C506.
Wistuba II, Behrens C, Milchgrub S, Syed S, Ahmadian M, Virmani AK et al. (1998). Clin Cancer Res 4: 2931–2938.
Zhao H, Jhanwar-Uniyal M, Datta PK, Yemul S, Ho L, Khitrov G et al. (2004). Int J Cancer 109: 65–70.
Acknowledgements
We are grateful to F Birg, D Maraninchi, C Mawas, A Puisieux and P Viens for discussions and encouragements. This work has been supported by Inserm, Institut Paoli-Calmettes, and grants from Ligue Nationale Contre le Cancer (Label) and Ministries of Health and Research (Cancéropôle PACA).
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on Oncogene website (http://www.nature.com/onc)
Rights and permissions
About this article
Cite this article
Charafe-Jauffret, E., Ginestier, C., Monville, F. et al. Gene expression profiling of breast cell lines identifies potential new basal markers. Oncogene 25, 2273–2284 (2006). https://doi.org/10.1038/sj.onc.1209254
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1209254
Keywords
This article is cited by
-
De-differentiation in cultures of organoids from luminal-type breast cancer is restored by inhibition of NOTCH signaling
Human Cell (2023)
-
Andrographolide elevates tumor necrosis factor-related apoptosis-inducing ligand lethality through reactive oxygen species accumulation and gasdermin E cleavage in breast cancer cells
Medical Oncology (2022)
-
Glycosphingolipids in human embryonic stem cells and breast cancer stem cells, and potential cancer therapy strategies based on their structures and functions
Glycoconjugate Journal (2022)
-
Hepatocyte growth factor pathway expression in breast cancer by race and subtype
Breast Cancer Research (2021)
-
Secreted breast tumor interstitial fluid microRNAs and their target genes are associated with triple-negative breast cancer, tumor grade, and immune infiltration
Breast Cancer Research (2020)