Abstract
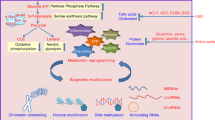
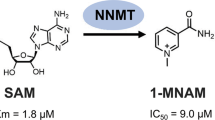
Nicotinamide N-methyltransferase (NNMT) is overexpressed in a variety of human cancers, where it contributes to tumorigenesis by a mechanism that is still poorly understood. Here we show using metabolomics that NNMT impairs the methylation potential of cancer cells by consuming methyl units from S-adenosyl methionine to create the stable metabolic product 1-methylnicotinamide. As a result, NNMT-expressing cancer cells have an altered epigenetic state that includes hypomethylated histones and other cancer-related proteins combined with heightened expression of protumorigenic gene products. Our findings thus point to a direct mechanistic link between the deregulation of a metabolic enzyme and widespread changes in the methylation landscape of cancer cells.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Warburg, O. On the origin of cancer cells. Science 123, 309–314 (1956).
DeBerardinis, R.J., Lum, J.J., Hatzivassiliou, G. & Thompson, C.B. The biology of cancer: metabolic reprogramming fuels cell growth and proliferation. Cell Metab. 7, 11–20 (2008).
Hsu, P.P. & Sabatini, D.M. Cancer cell metabolism: Warburg and beyond. Cell 134, 703–707 (2008).
Hanahan, D. & Weinberg, R.A. Hallmarks of cancer: the next generation. Cell 144, 646–674 (2011).
Wellen, K.E. & Thompson, C.B. A two-way street: reciprocal regulation of metabolism and signalling. Nat. Rev. Mol. Cell Biol. 13, 270–276 (2012).
DeBerardinis, R.J., Sayed, N., Ditsworth, D. & Thompson, C.B. Brick by brick: metabolism and tumor cell growth. Curr. Opin. Genet. Dev. 18, 54–61 (2008).
Possemato, R. et al. Functional genomics reveal that the serine synthesis pathway is essential in breast cancer. Nature 476, 346–350 (2011).
Nomura, D.K. et al. Monoacylglycerol lipase exerts dual control over endocannabinoid and fatty acid pathways to support prostate cancer. Chem. Biol. 18, 846–856 (2011).
Nomura, D.K., Dix, M.M. & Cravatt, B.F. Activity-based protein profiling for biochemical pathway discovery in cancer. Nat. Rev. Cancer 10, 630–638 (2010).
Roessler, M. et al. Identification of nicotinamide N-methyltransferase as a novel serum tumor marker for colorectal cancer. Clin. Cancer Res. 11, 6550–6557 (2005).
Wu, Y., Siadaty, M.S., Berens, M.E., Hampton, G.M. & Theodorescu, D. Overlapping gene expression profiles of cell migration and tumor invasion in human bladder cancer identify metallothionein 1E and nicotinamide N-methyltransferase as novel regulators of cell migration. Oncogene 27, 6679–6689 (2008).
Kim, J. et al. Expression of nicotinamide N-methyltransferase in hepatocellular carcinoma is associated with poor prognosis. J. Exp. Clin. Cancer Res. 28, 20 (2009).
Tomida, M., Mikami, I., Takeuchi, S., Nishimura, H. & Akiyama, H. Serum levels of nicotinamide N-methyltransferase in patients with lung cancer. J. Cancer Res. Clin. Oncol. 135, 1223–1229 (2009).
Tang, S.-W. Nicotinamide N-methyltransferase induces cellular invasion through activating matrix metalloproteinase-2 expression in clear cell renal cell carcinoma cells. Carcinogenesis 32, 138–145 (2011).
Nomura, D.K. et al. Monoacylglycerol lipase regulates a fatty acid network that promotes cancer pathogenesis. Cell 140, 49–61 (2010).
Peng, Y. et al. Structural basis of substrate recognition in human nicotinamide N-methyltransferase. Biochemistry 50, 7800–7808 (2011).
Bajad, S.U. et al. Separation and quantitation of water soluble cellular metabolites by hydrophilic interaction chromatography-tandem mass spectrometry. J. Chromatogr. A 1125, 76–88 (2006).
Saghatelian, A. et al. Assignment of endogenous substrates to enzymes by global metabolite profiling. Biochemistry 43, 14332–14339 (2004).
Smith, C.A., Want, E.J., O′Maille, G., Abagyan, R. & Siuzdak, G. XCMS:processing mass spectrometry data for metabolite profiling using nonlinear peak alignment, matching, and identification. Anal. Chem. 78, 779–787 (2006).
Zinellu, A. et al. Plasma methionine determination by capillary electrophoresis–UV assay: application on patients affected by retinal venous occlusive disease. Anal. Biochem. 363, 91–96 (2007).
Lu, S.C. S-Adenosylmethionine. Int. J. Biochem. Cell Biol. 32, 391–395 (2000).
Ulrey, C.L., Liu, L., Andrews, L.G. & Tollefsbol, T.O. The impact of metabolism on DNA methylation. Hum. Mol. Genet. 14, R139–R147 (2005).
Jia, G. et al. N6-Methyladenosine in nuclear RNA is a major substrate of the obesity-associated FTO. Nat. Chem. Biol. 7, 885–887 (2011).
Luo, M. Current chemical biology approaches to interrogate protein methyltransferases. ACS Chem. Biol. 7, 443–463 (2012).
Cantoni, G.L. The role of S-adenosylhomocysteine in the biological utilization of S-adenosylmethionine. Prog. Clin. Biol. Res. 198, 47–65 (1985).
Dawson, M.A. & Kouzarides, T. Cancer epigenetics: from mechanism to therapy. Cell 150, 12–27 (2012).
Varier, R.A. & Timmers, H.T. Histone lysine methylation and demethylation pathways in cancer. Biochim. Biophys. Acta 1815, 75–89 (2011).
Eichhorn, P.J.A., Creyghton, M.P. & Bernards, R. Protein phosphatase 2A regulatory subunits and cancer. Biochim. Biophys. Acta 1795, 1–15 (2009).
Puustinen, P. et al. PME-1 protects extracellular signal-regulated kinase pathway activity from protein phosphatase 2A–mediated inactivation in human malignant glioma. Cancer Res. 69, 2870–2877 (2009).
Boisvert, F.-M., Côté, J., Boulanger, M.-C. & Richard, S. A proteomic analysis of arginine-methylated protein complexes. Mol. Cell. Proteomics 2, 1319–1330 (2003).
Yang, H.W., Menon, L., Black, P., Carroll, R. & Johnson, M. SNAI2/Slug promotes growth and invasion in human gliomas. BMC Cancer 10, 301 (2010).
Zavadil, J. & Böttinger, E.P. TGF-β and epithelial-to-mesenchymal transitions. Oncogene 24, 5764–5774 (2005).
Su, J.-L. Knockdown of contactin-1 expression suppresses invasion and metastasis of lung adenocarcinoma. Cancer Res. 66, 2553–2561 (2006).
Wierinckx, A. et al. A diagnostic marker set for invasion, proliferation, and aggressiveness of prolactin pituitary tumors. Endocr. Relat. Cancer 14, 887–900 (2007).
Miyazaki, K. Laminin-5 (laminin-332): unique biological activity and role in tumor growth and invasion. Cancer Sci. 97, 91–98 (2006).
Dreger, H. et al. Epigenetic regulation of cell adhesion and communication by enhancer of zeste homolog 2 in human endothelial cells. Hypertension 60, 1176–1183 (2012).
Tan, J. et al. Pharmacologic disruption of Polycomb-repressive complex 2–mediated gene repression selectively induces apoptosis in cancer cells. Genes Dev. 21, 1050–1063 (2007).
Miranda, T.B. et al. DZNep is a global histone methylation inhibitor that reactivates developmental genes not silenced by DNA methylation. Mol. Cancer Ther. 8, 1579–1588 (2009).
Turcan, S. et al. IDH1 mutation is sufficient to establish the glioma hypermethylator phenotype. Nature 483, 479–483 (2012).
Dang, L. et al. Cancer-associated IDH1 mutations produce 2-hydroxyglutarate. Nature 462, 739–744 (2009).
Xu, W. et al. Oncometabolite 2-hydroxyglutarate is a competitive inhibitor of α-ketoglutarate–dependent dioxygenases. Cancer Cell 19, 17–30 (2011).
Wang, Y.C., Tang, F.Y., Chen, S.Y., Chen, Y.M. & Chiang, E.P. Glycine-N-methyltransferase expression in HepG2 cells is involved in methyl group homeostasis by regulating transmethylation kinetics and DNA methylation. J. Nutr. 141, 777–782 (2011).
Varela-Rey, M. et al. Fatty liver and fibrosis in glycine N-methyltransferase knockout mice is prevented by nicotinamide. Hepatology 52, 105–114 (2010).
Shyh-Chang, N. et al. Influence of threonine metabolism on S-adenosylmethionine and histone methylation. Science 339, 222–226 (2013).
Bitterman, K.J., Anderson, R.M., Cohen, H.Y., Latorre-Esteves, M. & Sinclair, D.A. Inhibition of silencing and accelerated aging by nicotinamide, a putative negative regulator of yeast sir2 and human SIRT1. J. Biol. Chem. 277, 45099–45107 (2002).
Smythe, G.A. et al. Concurrent quantification of quinolinic, picolinic, and nicotinic acids using electron-capture negative-ion gas chromatography-mass spectrometry. Anal. Biochem. 301, 21–26 (2002).
Catz, P. et al. Simultaneous determination of myristyl nicotinate, nicotinic acid, and nicotinamide in rabbit plasma by liquid chromatography-tandem mass spectrometry using methyl ethyl ketone as a deproteinization solvent. J. Chromatogr. B Analyt. Technol. Biomed. Life Sci. 829, 123–135 (2005).
Wardman, P. Chemical radiosensitizers for use in radiotherapy. Clin. Oncol. (R. Coll. Radiol.) 19, 397–417 (2007).
Joyce, T., Cantarella, D., Isella, C., Medico, E. & Pintzas, A. A molecular signature for epithelial to mesenchymal transition in a human colon cancer cell system is revealed by large-scale microarray analysis. Clin. Exp. Metastasis 26, 569–587 (2009).
Tomida, M., Ohtake, H., Yokota, T., Kobayashi, Y. & Kurosumi, M. Stat3 up-regulates expression of nicotinamide N-methyltransferase in human cancer cells. J. Cancer Res. Clin. Oncol. 134, 551–559 (2008).
French, J.B. et al. Characterization of nicotinamidases: steady state kinetic parameters, classwide inhibition by nicotinaldehydes, and catalytic mechanism. Biochemistry 49, 10421–10439 (2010).
Polkowska, J. et al. A combined experimental and theoretical study of the pH-dependent binding mode of NAD+ by water-soluble molecular clips. J. Phys. Org. Chem. 22, 779–790 (2009).
Martin, B.R., Giepmans, B.N., Adams, S.R. & Tsien, R.Y. Mammalian cell–based optimization of the biarsenical-binding tetracysteine motif for improved fluorescence and affinity. Nat. Biotechnol. 23, 1308–1314 (2005).
Yuan, J., Bennett, B.D. & Rabinowitz, J.D. Kinetic flux profiling for quantitation of cellular metabolic fluxes. Nat. Protoc. 3, 1328–1340 (2008).
Song, L., James, S.R., Kazim, L. & Karpf, A.R. Specific method for the determination of genomic DNA methylation by liquid chromatography-electrospray ionization tandem mass spectrometry. Anal. Chem. 77, 504–510 (2005).
Acknowledgements
This work was supported by the US National Institutes of Health (CA132630), a postdoctoral fellowship from Bayer (O.A.U.) and an US National Science Foundation predoctoral fellowship (A.M.Z.).
Author information
Authors and Affiliations
Contributions
O.A.U. and B.F.C. designed the experiments, analyzed the data and wrote the manuscript. A.M.Z. synthesized 1MNA and d4-1MNA. O.A.U. performed all other experiments.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Results (PDF 10684 kb)
Supplementary Data Set 1
Supplementary Dataset 1 (XLSX 0 kb)
Supplementary Data Set 2
Supplementary Dataset 2 (XLSX 2316 kb)
Rights and permissions
About this article
Cite this article
Ulanovskaya, O., Zuhl, A. & Cravatt, B. NNMT promotes epigenetic remodeling in cancer by creating a metabolic methylation sink. Nat Chem Biol 9, 300–306 (2013). https://doi.org/10.1038/nchembio.1204
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.1204
This article is cited by
-
Knockdown of nicotinamide N-methyltransferase suppresses proliferation, migration, and chemoresistance of Merkel cell carcinoma cells in vitro
Human Cell (2024)
-
Stromal nicotinamide N-methyltransferase orchestrates the crosstalk between fibroblasts and tumour cells in oral squamous cell carcinoma: evidence from patient-derived assembled organoids
Oncogene (2023)
-
Ligand-based in silico identification and biological evaluation of potential inhibitors of nicotinamide N-methyltransferase
Molecular Diversity (2023)
-
NNMT promotes the progression of intrahepatic cholangiocarcinoma by regulating aerobic glycolysis via the EGFR-STAT3 axis
Oncogenesis (2022)
-
The significance of NAD + metabolites and nicotinamide N-methyltransferase in chronic kidney disease
Scientific Reports (2022)